Chemical biology-whole genome engineering datasets predict new antibacterial combinations
- PMID: 34874820
- PMCID: PMC8767339
- DOI: 10.1099/mgen.0.000718
Chemical biology-whole genome engineering datasets predict new antibacterial combinations
Abstract
Trimethoprim and sulfamethoxazole are used commonly together as cotrimoxazole for the treatment of urinary tract and other infections. The evolution of resistance to these and other antibacterials threatens therapeutic options for clinicians. We generated and analysed a chemical-biology-whole-genome data set to predict new targets for antibacterial combinations with trimethoprim and sulfamethoxazole. For this we used a large transposon mutant library in Escherichia coli BW25113 where an outward-transcribing inducible promoter was engineered into one end of the transposon. This approach allows regulated expression of adjacent genes in addition to gene inactivation at transposon insertion sites, a methodology that has been called TraDIS-Xpress. These chemical genomic data sets identified mechanisms for both reduced and increased susceptibility to trimethoprim and sulfamethoxazole. The data identified that over-expression of FolA reduced trimethoprim susceptibility, a known mechanism for reduced susceptibility. In addition, transposon insertions into the genes tdk, deoR, ybbC, hha, ldcA, wbbK and waaS increased susceptibility to trimethoprim and likewise for rsmH, fadR, ddlB, nlpI and prc with sulfamethoxazole, while insertions in ispD, uspC, minC, minD, yebK, truD and umpG increased susceptibility to both these antibiotics. Two of these genes' products, Tdk and IspD, are inhibited by AZT and fosmidomycin respectively, antibiotics that are known to synergise with trimethoprim. Thus, the data identified two known targets and several new target candidates for the development of co-drugs that synergise with trimethoprim, sulfamethoxazole or cotrimoxazole. We demonstrate that the TraDIS-Xpress technology can be used to generate information-rich chemical-genomic data sets that can be used for antibacterial development.
Keywords: TIS; TraDIS; antibiotics; chemical; combinations; genomics.
Conflict of interest statement
No authors have any conflicts of interest to declare. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Figures


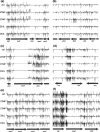

References
-
- Gangcuangco LM, Alejandria M, Henson KE, Alfaraz L, Ata RM, et al. Prevalence and risk factors for trimethoprim-sulfamethoxazole-resistant Escherichia coli among women with acute uncomplicated urinary tract infection in a developing country. Int J Infect Dis. 2015;34:55–60. doi: 10.1016/j.ijid.2015.02.022. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

