Gene excavation and expression analysis of CYP and UGT related to the post modifying stage of gypenoside biosynthesis in Gynostemma pentaphyllum (Thunb.) Makino by comprehensive analysis of RNA and proteome sequencing
- PMID: 34874937
- PMCID: PMC8651138
- DOI: 10.1371/journal.pone.0260027
Gene excavation and expression analysis of CYP and UGT related to the post modifying stage of gypenoside biosynthesis in Gynostemma pentaphyllum (Thunb.) Makino by comprehensive analysis of RNA and proteome sequencing
Abstract
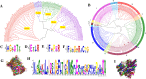
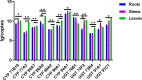
Previous studies have revealed that gypenosides produced from Gynostemma pentaphyllum (Thunb.) Makino are mainly dammarane-type triterpenoid saponins with diverse structures and important biological activities, but the mechanism of diversity for gypenoside biosynthesis is still unclear. In this study, a combination of isobaric tags for relative and absolute quantification (iTRAQ) proteome analysis and RNA sequencing transcriptome analysis was performed to identify the proteins and genes related to gypenoside biosynthesis. A total of 3925 proteins were identified by proteomic sequencing, of which 2537 were quantified. Seventeen cytochrome P450 (CYP) and 11 uridine 5'-diphospho-glucuronosyltransferase (UDP-glucuronosyltransferase, UGT) candidate genes involved in the side chain synthesis and modification of gypenosides were found. Seven putative CYPs (CYP71B19, CYP77A3, CYP86A7, CYP86A8, CYP89A2, CYP90A1, CYP94A1) and five putative UGTs (UGT73B4, UGT76B1, UGT74F2, UGT91C1 and UGT91A1) were selected as candidate structural modifiers of triterpenoid saponins, which were cloned for gene expression analysis. Comprehensive analysis of RNA sequencing and proteome sequencing showed that some CYPs and UGTs were found at both the transcription and translation levels. In this study, an expression analysis of 7 CYPs and 5 UGTs that contributed to gypenoside biosynthesis and distribution in G. pentaphyllum was performed, providing consistent results that will inspire more future research on vital genes/proteins involved in gypenoside biosynthesis.
Conflict of interest statement
No authors have competing interests.
Figures


References
-
- Chen L, Brar MS, Leung FCC, Hsiao WLW. Triterpenoid herbal saponins enhance beneficial bacteria, decrease sulfate-reducing bacteria, modulate inflammatory intestinal microenvironment and exert cancer preventive effects in ApcMin/+ mice. Oncotarget. 2016;7(21):31226–42. doi: 10.18632/oncotarget.8886 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

