eIF2B-capturing viral protein NSs suppresses the integrated stress response
- PMID: 34876589
- PMCID: PMC8651795
- DOI: 10.1038/s41467-021-27337-x
eIF2B-capturing viral protein NSs suppresses the integrated stress response
Abstract
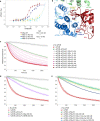
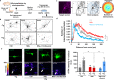
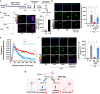
Various stressors such as viral infection lead to the suppression of cap-dependent translation and the activation of the integrated stress response (ISR), since the stress-induced phosphorylated eukaryotic translation initiation factor 2 [eIF2(αP)] tightly binds to eIF2B to prevent it from exchanging guanine nucleotide molecules on its substrate, unphosphorylated eIF2. Sandfly fever Sicilian virus (SFSV) evades this cap-dependent translation suppression through the interaction between its nonstructural protein NSs and host eIF2B. However, its precise mechanism has remained unclear. Here, our cryo-electron microscopy (cryo-EM) analysis reveals that SFSV NSs binds to the α-subunit of eIF2B in a competitive manner with eIF2(αP). Together with SFSV NSs, eIF2B retains nucleotide exchange activity even in the presence of eIF2(αP), in line with the cryo-EM structures of the eIF2B•SFSV NSs•unphosphorylated eIF2 complex. A genome-wide ribosome profiling analysis clarified that SFSV NSs expressed in cultured human cells attenuates the ISR triggered by thapsigargin, an endoplasmic reticulum stress inducer. Furthermore, SFSV NSs introduced in rat hippocampal neurons and human induced-pluripotent stem (iPS) cell-derived motor neurons exhibits neuroprotective effects against the ISR-inducing stress. Since ISR inhibition is beneficial in various neurological disease models, SFSV NSs may be a promising therapeutic ISR inhibitor.
© 2021. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures
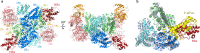


Similar articles
-
Viral evasion of the integrated stress response through antagonism of eIF2-P binding to eIF2B.Nat Commun. 2021 Dec 7;12(1):7103. doi: 10.1038/s41467-021-26164-4. Nat Commun. 2021. PMID: 34876554 Free PMC article.
-
Protection of eIF2B from inhibitory phosphorylated eIF2: A viral strategy to maintain mRNA translation during the PKR-triggered integrated stress response.J Biol Chem. 2023 Nov;299(11):105287. doi: 10.1016/j.jbc.2023.105287. Epub 2023 Sep 22. J Biol Chem. 2023. PMID: 37742919 Free PMC article. Review.
-
eIF2B as a Target for Viral Evasion of PKR-Mediated Translation Inhibition.mBio. 2020 Jul 14;11(4):e00976-20. doi: 10.1128/mBio.00976-20. mBio. 2020. PMID: 32665273 Free PMC article.
-
ISRIB Blunts the Integrated Stress Response by Allosterically Antagonising the Inhibitory Effect of Phosphorylated eIF2 on eIF2B.Mol Cell. 2021 Jan 7;81(1):88-103.e6. doi: 10.1016/j.molcel.2020.10.031. Epub 2020 Nov 20. Mol Cell. 2021. PMID: 33220178 Free PMC article.
-
Crystal structure of eIF2B and insights into eIF2-eIF2B interactions.FEBS J. 2017 Mar;284(6):868-874. doi: 10.1111/febs.13896. Epub 2016 Sep 29. FEBS J. 2017. PMID: 27627185 Review.
Cited by
-
dCas13-mediated translational repression for accurate gene silencing in mammalian cells.Nat Commun. 2024 Mar 11;15(1):2205. doi: 10.1038/s41467-024-46412-7. Nat Commun. 2024. PMID: 38467613 Free PMC article.
-
Selection of initiator tRNA and start codon by mammalian mitochondrial initiation factor 3 in leaderless mRNA translation.Nucleic Acids Res. 2025 Jan 24;53(3):gkaf021. doi: 10.1093/nar/gkaf021. Nucleic Acids Res. 2025. PMID: 39878211 Free PMC article.
-
Turnip mosaic virus NIb weakens the function of eukaryotic translation initiation factor 6 facilitating viral infection in Nicotiana benthamiana.Mol Plant Pathol. 2024 Feb;25(2):e13434. doi: 10.1111/mpp.13434. Mol Plant Pathol. 2024. PMID: 38388027 Free PMC article.
-
TRMT10A dysfunction perturbs codon translation of initiator methionine and glutamine and impairs brain functions in mice.Nucleic Acids Res. 2024 Aug 27;52(15):9230-9246. doi: 10.1093/nar/gkae520. Nucleic Acids Res. 2024. PMID: 38950903 Free PMC article.
-
Mettl1-dependent m7G tRNA modification is essential for maintaining spermatogenesis and fertility in Drosophila melanogaster.Nat Commun. 2024 Sep 24;15(1):8147. doi: 10.1038/s41467-024-52389-0. Nat Commun. 2024. PMID: 39317727 Free PMC article.
References
-
- Kashiwagi K, et al. Structural basis for eIF2B inhibition in integrated stress response. Science. 2019;364:495–499. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

