Superhuman cell death detection with biomarker-optimized neural networks
- PMID: 34878844
- PMCID: PMC8654296
- DOI: 10.1126/sciadv.abf8142
Superhuman cell death detection with biomarker-optimized neural networks
Abstract
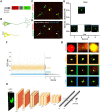
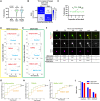
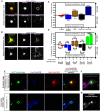
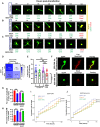
Cellular events underlying neurodegenerative disease may be captured by longitudinal live microscopy of neurons. While the advent of robot-assisted microscopy has helped scale such efforts to high-throughput regimes with the statistical power to detect transient events, time-intensive human annotation is required. We addressed this fundamental limitation with biomarker-optimized convolutional neural networks (BO-CNNs): interpretable computer vision models trained directly on biosensor activity. We demonstrate the ability of BO-CNNs to detect cell death, which is typically measured by trained annotators. BO-CNNs detected cell death with superhuman accuracy and speed by learning to identify subcellular morphology associated with cell vitality, despite receiving no explicit supervision to rely on these features. These models also revealed an intranuclear morphology signal that is difficult to spot by eye and had not previously been linked to cell death, but that reliably indicates death. BO-CNNs are broadly useful for analyzing live microscopy and essential for interpreting high-throughput experiments.
Figures



References
-
- Arrasate M., Mitra S., Schweitzer E. S., Segal M. R., Finkbeiner S., Inclusion body formation reduces levels of mutant huntingtin and the risk of neuronal death. Nature 431, 805–810 (2004). - PubMed
-
- Miller J., Arrasate M., Shaby B. A., Mitra S., Masliah E., Finkbeiner S., Quantitative relationships between huntingtin levels, polyglutamine length, inclusion body formation, and neuronal death provide novel insight into Huntington's disease molecular pathogenesis. J. Neurosci. 30, 10541–10550 (2010). - PMC - PubMed
-
- Carpenter A. E., Extracting rich information from images. Methods Mol. Biol. 486, 193–211 (2009). - PubMed

