Evaluation of Deep Neural Network ProSPr for Accurate Protein Distance Predictions on CASP14 Targets
- PMID: 34884640
- PMCID: PMC8657919
- DOI: 10.3390/ijms222312835
Evaluation of Deep Neural Network ProSPr for Accurate Protein Distance Predictions on CASP14 Targets
Abstract
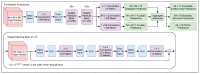
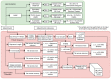
The field of protein structure prediction has recently been revolutionized through the introduction of deep learning. The current state-of-the-art tool AlphaFold2 can predict highly accurate structures; however, it has a prohibitively long inference time for applications that require the folding of hundreds of sequences. The prediction of protein structure annotations, such as amino acid distances, can be achieved at a higher speed with existing tools, such as the ProSPr network. Here, we report on important updates to the ProSPr network, its performance in the recent Critical Assessment of Techniques for Protein Structure Prediction (CASP14) competition, and an evaluation of its accuracy dependency on sequence length and multiple sequence alignment depth. We also provide a detailed description of the architecture and the training process, accompanied by reusable code. This work is anticipated to provide a solid foundation for the further development of protein distance prediction tools.
Keywords: CASP; ProSPr; alphafold; contact; dataset; deep learning; distance; prediction; protein; retrainable.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

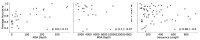
References
-
- Morris C.J., Corte D.D. Using molecular docking and molecular dynamics to investigate protein-ligand interactions. Mod. Phys. Lett. B. 2021;35:2130002. doi: 10.1142/S0217984921300027. - DOI
-
- Coates T.L., Young N., Jarrett A.J., Morris C.J., Moody J.D., Corte D.D. Current computational methods for enzyme design. Mod. Phys. Lett. B. 2021;35:2150155. doi: 10.1142/S0217984921501554. - DOI
-
- Möckel C., Kubiak J., Schillinger O., Kühnemuth R., Della Corte D., Schröder G.F., Willbold D., Strodel B., Seidel C.A., Neudecker P. Integrated NMR, fluorescence, and molecular dynamics benchmark study of protein mechanics and hydrodynamics. J. Phys. Chem. B. 2018;123:1453–1480. doi: 10.1021/acs.jpcb.8b08903. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

