Interaction of Klebsiella pneumoniae with tissue macrophages in a mouse infection model and ex-vivo pig organ perfusions: an exploratory investigation
- PMID: 34901898
- PMCID: PMC8641047
- DOI: 10.1016/S2666-5247(21)00195-6
Interaction of Klebsiella pneumoniae with tissue macrophages in a mouse infection model and ex-vivo pig organ perfusions: an exploratory investigation
Abstract
Background: Hypervirulent Klebsiella pneumoniae (hvKp) strains of capsule type K1 and K2 cause invasive infections associated with hepatic abscesses, which can be difficult to treat and are frequently associated with relapsing infections. Other K pneumoniae strains (non-hvKp), including lineages that have acquired carbapenem resistance, do not manifest this pathology. In this work we aimed to test the hypothesis that within-macrophage replication is a key mechanism underpinning abscess formation in hvKp infections.
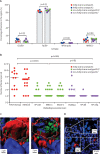
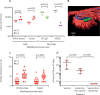
Methods: In this exploratory investigation, to study the pathophysiology of abscess formation, mice were intravenously infected with 106 colony forming units (CFU) of either hvKp isolates (six strains) or non-hvKp isolates (seven strains). Intracellular bacterial replication and neutrophil influx in liver and spleen was quantified by fluorescence microscopy of sliced cryopreserved organs of mice collected 30 min, 6 h, and 24 h after infection with the aim to provide data of bacterial association to Kupffer cells in the liver and to the different tissue macrophages in the spleen. Microbiological and microscopy analysis of an ex-vivo model of pig liver and spleen infection were used to confirm within-macrophage replication. Pig organs were perfused with heparinised, autologous pig's blood and injected with 6·5 × 107 CFU of hvKp K2 sequence type 25 strain GMR151. Blood and tissue biopsies collected before infection and 30 min, 1 h, 2 h, 3 h, 4 h, and 5 h after infection were used to measure bacterial counts and to identify the subcellular localisation of bacteria by immunohistochemistry analysis.
Findings: We show that hvKp resisted phagocyte-mediated clearance and replicated in mouse liver macrophages to form clusters 6 h after infection, with a mean of 7·0 bacteria per Kupffer cell (SD 6·2); however, non-hvKp were efficiently cleared (mean 1·5 bacteria per cell [SD 1·1]). HvKp infection promoted neutrophil recruitment to sites of infection, which in the liver resulted in histopathological signs of abscess formation as early as 24 h post-infection. Experiments in pig organs which share a high functional and anatomical resemblance to human organs, provided strong evidence for the propensity of hvKp to replicate within the hepatic macrophages.
Interpretation: These findings show subversion of innate immune processes in the liver by K pneumoniae and resistance to Kupffer cell mediated clearance as an explanation for the propensity of hvKp strains to cause hepatic abscesses.
Funding: University of Oxford and a Royal Society Wolfson grant funded biosafety facility.
© 2021 The Author(s). Published by Elsevier Ltd. This is an Open Access article under the CC BY 4.0 license.
Conflict of interest statement
MP is a full-time employee of GlaxoSmithKline. GMR reports grants, personal fees, and non-financial support from Accelerate Diagnostics, Beckman Coulter, and Menarini; grants from Seegene, Arrow, Symcel, DID, Hain Lifescience, Meridian, SetLance, Qvella, Qlinea, Biomedical Service, and Quidel; personal fees from Becton Dickinson, Pfizer, Roche, Thermo Fisher, QPex, and Qiagen; grants and personal fees from bioMeriéux, Cepheid, MSD, Nordic Pharma, Shionogi, Zambon, and Angelini; personal fees from Venatorx; and income from selling strains to Venatorx, outside the submitted work. MRO reports a research contract with the University of Oxford, a studentship from the Medical Research Council (MRC), and grants from Royal Society Wolfson and the Biotechnology and Biological Sciences Research Council (BBSRC), during the conduct of the study; grants and a studentship from GlaxoSmithKline, outside the submitted work. All other authors declare no competing interests.
Figures




References
-
- Siu LK, Yeh KM, Lin JC, Fung CP, Chang FY. Klebsiella pneumoniae liver abscess: a new invasive syndrome. Lancet Infect Dis. 2012;12:881–887. - PubMed
-
- Liu YC, Cheng DL, Lin CL. Klebsiella pneumoniae liver abscess associated with septic endophthalmitis. Arch Intern Med. 1986;146:1913–1916. - PubMed
-
- Fazili T, Sharngoe C, Endy T, Kiska D, Javaid W, Polhemus M. Klebsiella pneumoniae liver abscess: an emerging disease. Am J Med Sci. 2016;351:297–304. - PubMed
-
- Nordmann P, Cuzon G, Naas T. The real threat of Klebsiella pneumoniae carbapenemase-producing bacteria. Lancet Infect Dis. 2009;9:228–236. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

