High-affinity, neutralizing antibodies to SARS-CoV-2 can be made without T follicular helper cells
- PMID: 34914544
- PMCID: PMC8977051
- DOI: 10.1126/sciimmunol.abl5652
High-affinity, neutralizing antibodies to SARS-CoV-2 can be made without T follicular helper cells
Abstract
T follicular helper (TFH) cells are the conventional drivers of protective, germinal center (GC)–based antiviral antibody responses. However, loss of TFH cells and GCs has been observed in patients with severe COVID-19. As T cell–B cell interactions and immunoglobulin class switching still occur in these patients, noncanonical pathways of antibody production may be operative during SARS-CoV-2 infection. We found that both TFH-dependent and -independent antibodies were induced against SARS-CoV-2 infection, SARS-CoV-2 vaccination, and influenza A virus infection. Although TFH-independent antibodies to SARS-CoV-2 had evidence of reduced somatic hypermutation, they were still high affinity, durable, and reactive against diverse spike-derived epitopes and were capable of neutralizing both homologous SARS-CoV-2 and the B.1.351 (beta) variant of concern. We found by epitope mapping and B cell receptor sequencing that TFH cells focused the B cell response, and therefore, in the absence of TFH cells, a more diverse clonal repertoire was maintained. These data support an alternative pathway for the induction of B cell responses during viral infection that enables effective, neutralizing antibody production to complement traditional GC-derived antibodies that might compensate for GCs damaged by viral inflammation.
Conflict of interest statement
Competing interests:
KK, JB, WAH, and JCS declare the following competing interests: ownership of stocks or shares at Serimmune, paid employment at Serimmune, and patent applications on behalf of Serimmune. Yale University (CBW) has a patent pending entitled “Compounds and Compositions for Treating, Ameliorating, and/or Preventing SARS-CoV-2 Infection and/or Complications Thereof.”
Figures


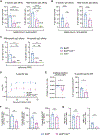




References
-
- McMahan K, Yu J, Mercado NB, Loos C, Tostanoski LH, Chandrashekar A, Liu J, Peter L, Atyeo C, Zhu A, Bondzie EA, Dagotto G, Gebre MS, Jacob-Dolan C, Li Z, Nampanya F, Patel S, Pessaint L, Van Ry A, Blade K, Yalley-Ogunro J, Cabus M, Brown R, Cook A, Teow E, Andersen H, Lewis MG, Lauffenburger DA, Alter G, Barouch DH, Correlates of protection against SARS-CoV-2 in rhesus macaques. Nature. 590, 630–634 (2021). - PMC - PubMed
-
- Zhang J, Wu Q, Liu Z, Wang Q, Wu J, Hu Y, Bai T, Xie T, Huang M, Wu T, Peng D, Huang W, Jin K, Niu L, Guo W, Luo D, Lei D, Wu Z, Li G, Huang R, Lin Y, Xie X, He S, Deng Y, Liu J, Li W, Lu Z, Chen H, Zeng T, Luo Q, Li Y-P, Wang Y, Liu W, Qu X, Spike-specific circulating T follicular helper cell and cross-neutralizing antibody responses in COVID-19-convalescent individuals. Nature Microbiology. 6, 51–58 (2021). - PubMed
-
- Moderbacher CR, Ramirez SI, Dan JM, Grifoni A, Hastie KM, Weiskopf D, Belanger S, Abbott RK, Kim C, Choi J, Kato Y, Crotty EG, Kim C, Rawlings SA, Mateus J, Tse LPV, Frazier A, Baric R, Peters B, Greenbaum J, Saphire EO, Smith DM, Sette A, Crotty S, Antigen-Specific Adaptive Immunity to SARS-CoV-2 in Acute COVID-19 and Associations with Age and Disease Severity. Cell. 183, 996–1012.e19 (2020). - PMC - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
Grants and funding
- T32 AI007517/AI/NIAID NIH HHS/United States
- R01 AR074545/AR/NIAMS NIH HHS/United States
- F30 HL149151/HL/NHLBI NIH HHS/United States
- R01 AI148467/AI/NIAID NIH HHS/United States
- R01 AI136942/AI/NIAID NIH HHS/United States
- F30 CA239444/CA/NCI NIH HHS/United States
- K08 AI128043/AI/NIAID NIH HHS/United States
- F30 CA250249/CA/NCI NIH HHS/United States
- T32 HL007974/HL/NHLBI NIH HHS/United States
- R37 AR040072/AR/NIAMS NIH HHS/United States
- T32 AI007019/AI/NIAID NIH HHS/United States
- UL1 TR001863/TR/NCATS NIH HHS/United States
- T32 GM136651/GM/NIGMS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

