An integrative bioinformatics analysis for identifying hub genes associated with infection of lung samples in patients infected with SARS-CoV-2
- PMID: 34920753
- PMCID: PMC8677925
- DOI: 10.1186/s40001-021-00609-4
An integrative bioinformatics analysis for identifying hub genes associated with infection of lung samples in patients infected with SARS-CoV-2
Abstract
Background: At the end of 2019, the world witnessed the emergence and ravages of a viral infection induced by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). Also known as the coronavirus disease 2019 (COVID-19), it has been identified as a public health emergency of international concern (PHEIC) by the World Health Organization (WHO) because of its severity.
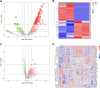
Methods: The gene data of 51 samples were extracted from the GSE150316 and GSE147507 data set and then processed by means of the programming language R, through which the differentially expressed genes (DEGs) that meet the standards were screened. The Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses were performed on the selected DEGs to understand the functions and approaches of DEGs. The online tool STRING was employed to construct a protein-protein interaction (PPI) network of DEGs and, in turn, to identify hub genes.
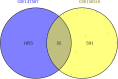
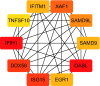
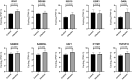
Results: A total of 52 intersection genes were obtained through DEG identification. Through the GO analysis, we realized that the biological processes (BPs) that have the deepest impact on the human body after SARS-CoV-2 infection are various immune responses. By using STRING to construct a PPI network, 10 hub genes were identified, including IFIH1, DDX58, ISG15, EGR1, OASL, SAMD9, SAMD9L, XAF1, IFITM1, and TNFSF10.
Conclusion: The results of this study will hopefully provide guidance for future studies on the pathophysiological mechanism of SARS-CoV-2 infection.
Keywords: Differentially expressed genes; Hub genes; Protein–protein interactions network; SARS-CoV-2.
© 2021. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures





Similar articles
-
Analysis and Identification Genetic Effect of SARS-CoV-2 Infections to Alzheimer's Disease Patients by Integrated Bioinformatics.J Alzheimers Dis. 2022;85(2):729-744. doi: 10.3233/JAD-215086. J Alzheimers Dis. 2022. PMID: 34776447
-
Exploration and validation of related hub gene expression during SARS-CoV-2 infection of human bronchial organoids.Hum Genomics. 2021 Mar 16;15(1):18. doi: 10.1186/s40246-021-00316-5. Hum Genomics. 2021. PMID: 33726831 Free PMC article.
-
Identification of the Hub Genes and the Signaling Pathways in Human iPSC-Cardiomyocytes Infected by SARS-CoV-2.Biochem Genet. 2022 Dec;60(6):2052-2068. doi: 10.1007/s10528-022-10206-7. Epub 2022 Mar 2. Biochem Genet. 2022. PMID: 35235083 Free PMC article.
-
Gene expression profiling of host lipid metabolism in SARS-CoV-2 infected patients: a systematic review and integrated bioinformatics analysis.BMC Infect Dis. 2024 Jan 23;24(1):124. doi: 10.1186/s12879-024-08983-0. BMC Infect Dis. 2024. PMID: 38263024 Free PMC article.
-
Hidden in plain sight: The effects of BCG vaccination in the COVID-19 pandemic.J Med Virol. 2021 Apr;93(4):1950-1966. doi: 10.1002/jmv.26707. Epub 2020 Dec 17. J Med Virol. 2021. PMID: 33289122 Free PMC article. Review.
Cited by
-
Role of early growth response 1 in inflammation-associated lung diseases.Am J Physiol Lung Cell Mol Physiol. 2023 Aug 1;325(2):L143-L154. doi: 10.1152/ajplung.00413.2022. Epub 2023 Jul 4. Am J Physiol Lung Cell Mol Physiol. 2023. PMID: 37401387 Free PMC article. Review.
-
Examining the role of EGR1 during viral infections.Front Microbiol. 2022 Oct 21;13:1020220. doi: 10.3389/fmicb.2022.1020220. eCollection 2022. Front Microbiol. 2022. PMID: 36338037 Free PMC article. Review.
-
Five-Year Assessment of Multiple Gene Variants Associated with Bone Marrow Hypocellularity, Reduced Bone Density, and Ovarian Insufficiency in Adolescence.J Bone Metab. 2022 Nov;29(4):271-277. doi: 10.11005/jbm.2022.29.4.271. Epub 2022 Nov 30. J Bone Metab. 2022. PMID: 36529870 Free PMC article.
-
Immune-Related Protein Interaction Network in Severe COVID-19 Patients toward the Identification of Key Proteins and Drug Repurposing.Biomolecules. 2022 May 11;12(5):690. doi: 10.3390/biom12050690. Biomolecules. 2022. PMID: 35625619 Free PMC article.
References
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

