DNA damage checkpoint and repair: From the budding yeast Saccharomyces cerevisiae to the pathogenic fungus Candida albicans
- PMID: 34938410
- PMCID: PMC8645783
- DOI: 10.1016/j.csbj.2021.11.033
DNA damage checkpoint and repair: From the budding yeast Saccharomyces cerevisiae to the pathogenic fungus Candida albicans
Abstract
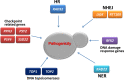
Cells are constantly challenged by internal or external genotoxic assaults, which may induce a high frequency of DNA lesions, leading to genome instability. Accumulation of damaged DNA is severe or even lethal to cells and can result in abnormal proliferation that can cause cancer in multicellular organisms, aging or cell death. Eukaryotic cells have evolved a comprehensive defence system termed the DNA damage response (DDR) to monitor and remove lesions in their DNA. The DDR has been extensively studied in the budding yeast Saccharomyces cerevisiae. Emerging evidence indicates that DDR genes in the pathogenic fungus Candida albicans show functional consistency with their orthologs in S. cerevisiae, but may act through distinct mechanisms. In particular, the DDR in C. albicans appears critical for resisting DNA damage stress induced by reactive oxygen species (ROS) produced from immune cells, and this plays a vital role in pathogenicity. Therefore, DDR genes could be considered as potential targets for clinical therapies. This review summarizes the identified DNA damage checkpoint and repair genes in C. albicans based on their orthologs in S. cerevisiae, and discusses their contribution to pathogenicity in C. albicans.
Keywords: Candida albicans; DNA damage checkpoint; DNA damage repair; DNA damage response; Pathogenicity.
© 2021 The Authors.
Conflict of interest statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures


Similar articles
-
Nucleotide Excision Repair Protein Rad23 Regulates Cell Virulence Independent of Rad4 in Candida albicans.mSphere. 2020 Feb 19;5(1):e00062-20. doi: 10.1128/mSphere.00062-20. mSphere. 2020. PMID: 32075883 Free PMC article.
-
Transcriptional Profiling of the Candida albicans Response to the DNA Damage Agent Methyl Methanesulfonate.Int J Mol Sci. 2022 Jul 7;23(14):7555. doi: 10.3390/ijms23147555. Int J Mol Sci. 2022. PMID: 35886903 Free PMC article.
-
The Role of Mms22p in DNA Damage Response in Candida albicans.G3 (Bethesda). 2015 Oct 4;5(12):2567-78. doi: 10.1534/g3.115.021840. G3 (Bethesda). 2015. PMID: 26438292 Free PMC article.
-
How do cells sense DNA lesions?Biochem Soc Trans. 2020 Apr 29;48(2):677-691. doi: 10.1042/BST20191118. Biochem Soc Trans. 2020. PMID: 32219379 Review.
-
Yeast signaling pathways in the oxidative stress response.Mutat Res. 2005 Jan 6;569(1-2):13-27. doi: 10.1016/j.mrfmmm.2004.09.006. Mutat Res. 2005. PMID: 15603750 Review.
Cited by
-
Importance of the Aspergillus fumigatus Mismatch Repair Protein Msh6 in Antifungal Resistance Development.J Fungi (Basel). 2024 Mar 12;10(3):210. doi: 10.3390/jof10030210. J Fungi (Basel). 2024. PMID: 38535218 Free PMC article.
-
DNA Damage Checkpoints Govern Global Gene Transcription and Exhibit Species-Specific Regulation on HOF1 in Candida albicans.J Fungi (Basel). 2024 May 29;10(6):387. doi: 10.3390/jof10060387. J Fungi (Basel). 2024. PMID: 38921373 Free PMC article.
-
Enhanced multistress tolerance of Saccharomyces cerevisiae with the sugar transporter-like protein Stl1F427L mutation in the presence of glycerol.Microbiol Spectr. 2025 Feb 4;13(2):e0008924. doi: 10.1128/spectrum.00089-24. Epub 2024 Dec 16. Microbiol Spectr. 2025. PMID: 39679667 Free PMC article.
-
DNA damage repair factor Rad18 controls virulence partially via transcriptional suppression of genes HWP1 and ECE1 in Candida albicans.Virulence. 2024 Dec;15(1):2433201. doi: 10.1080/21505594.2024.2433201. Epub 2024 Nov 27. Virulence. 2024. PMID: 39573927 Free PMC article.
-
Mec1-Rad53 Signaling Regulates DNA Damage-Induced Autophagy and Pathogenicity in Candida albicans.J Fungi (Basel). 2023 Dec 9;9(12):1181. doi: 10.3390/jof9121181. J Fungi (Basel). 2023. PMID: 38132782 Free PMC article.
References
-
- Papon N., Courdavault V., Clastre M., Bennett R.J., Heitman J. Emerging and emerged pathogenic Candida species: beyond the Candida albicans paradigm. PLoS Pathog. 2013;9(9):e1003550. doi: 10.1371/journal.ppat.100355010.1371/journal.ppat.1003550.g00110.1371/journal.ppat.1003550.t001. - DOI - PMC - PubMed
Publication types
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
