Cervical Cancer Development: Implications of HPV16 E6E7-NFX1-123 Regulated Genes
- PMID: 34944802
- PMCID: PMC8699269
- DOI: 10.3390/cancers13246182
Cervical Cancer Development: Implications of HPV16 E6E7-NFX1-123 Regulated Genes
Abstract
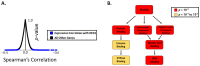
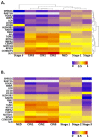
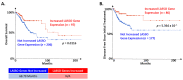
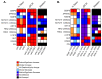
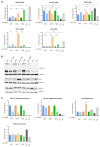
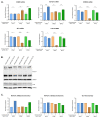
High-risk human papillomavirus (HR HPV) causes nearly all cervical cancers, half of which are due to HPV type 16 (HPV16). HPV16 oncoprotein E6 (16E6) binds to NFX1-123, and dysregulates gene expression, but their clinical implications are unknown. Additionally, HPV16 E7's role has not been studied in concert with NFX1-123 and 16E6. HR HPVs express both oncogenes, and transformation requires their expression, so we sought to investigate the effect of E7 on gene expression. This study's goal was to define gene expression profiles across cervical precancer and cancer stages, identify genes correlating with disease progression, assess patient survival, and validate findings in cell models. We analyzed NCBI GEO datasets containing transcriptomic data linked with cervical cancer stage and utilized LASSO analysis to identify cancer-driving genes. Keratinocytes expressing 16E6 and 16E7 (16E6E7) and exogenous NFX1-123 were tested for LASSO-identified gene expression. Ten out of nineteen genes correlated with disease progression, including CEBPD, NOTCH1, and KRT16, and affected survival. 16E6E7 in keratinocytes increased CEBPD, KRT16, and SLPI, and decreased NOTCH1. Exogenous NFX1-123 in 16E6E7 keratinocytes resulted in significantly increased CEBPD and NOTCH1, and reduced SLPI. This work demonstrates the clinical relevance of CEBPD, NOTCH1, KRT16, and SLPI, and shows the regulatory effects of 16E6E7 and NFX1-123.
Keywords: E6; E7; HPV16; NFX1-123; cervical cancer; keratinocytes.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

References
-
- Brianti P., De Flammineis E., Mercuri S.R. Review of HPV-related diseases and cancers. New Microbiol. 2017;40:80–85. - PubMed
-
- Giuliano A.R., Lee J.H., Fulp W., Villa L.L., Lazcano E., Papenfuss M.R., Abrahamsen M., Salmeron J., Anic G.M., Rollison D.E., et al. Incidence and clearance of genital human papillomavirus infection in men (HIM): A cohort study. Lancet. 2011;377:932–940. doi: 10.1016/S0140-6736(10)62342-2. - DOI - PMC - PubMed
-
- Kreimer A.R., Pierce Campbell C.M., Lin H.Y., Fulp W., Papenfuss M.R., Abrahamsen M., Hildesheim A., Villa L.L., Salmeron J.J., Lazcano-Ponce E., et al. Incidence and clearance of oral human papillomavirus infection in men: The HIM cohort study. Lancet. 2013;382:877–887. doi: 10.1016/S0140-6736(13)60809-0. - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

