Arginine and Arginases Modulate Metabolism, Tumor Microenvironment and Prostate Cancer Progression
- PMID: 34960055
- PMCID: PMC8704013
- DOI: 10.3390/nu13124503
Arginine and Arginases Modulate Metabolism, Tumor Microenvironment and Prostate Cancer Progression
Abstract
Arginine availability and activation of arginine-related pathways at cancer sites have profound effects on the tumor microenvironment, far beyond their well-known role in the hepatic urea cycle. Arginine metabolism impacts not only malignant cells but also the surrounding immune cells behavior, modulating growth, survival, and immunosurveillance mechanisms, either through an arginase-mediated effect on polyamines and proline synthesis, or by the arginine/nitric oxide pathway in tumor cells, antitumor T-cells, myeloid-derived suppressor cells, and macrophages. This review presents evidence concerning the impact of arginine metabolism and arginase activity in the prostate cancer microenvironment, highlighting the recent advances in immunotherapy, which might be relevant for prostate cancer. Even though further research is required, arginine deprivation may represent a novel antimetabolite strategy for the treatment of arginine-dependent prostate cancer.
Keywords: arginase; arginine; metabolism; nitric oxide; prostate cancer; tumor microenvironment.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
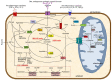


Similar articles
-
Metabolomic reprogramming of the tumor microenvironment by dual arginase inhibitor OATD-02 boosts anticancer immunity.Sci Rep. 2025 May 28;15(1):18741. doi: 10.1038/s41598-025-03446-1. Sci Rep. 2025. PMID: 40437024 Free PMC article.
-
Myeloid Cell-Derived Arginase in Cancer Immune Response.Front Immunol. 2020 May 15;11:938. doi: 10.3389/fimmu.2020.00938. eCollection 2020. Front Immunol. 2020. PMID: 32499785 Free PMC article. Review.
-
Arginine availability, arginase, and the immune response.Curr Opin Clin Nutr Metab Care. 2003 Mar;6(2):223-8. doi: 10.1097/00075197-200303000-00012. Curr Opin Clin Nutr Metab Care. 2003. PMID: 12589193 Review.
-
Engineered microparticles modulate arginine metabolism to repolarize tumor-associated macrophages for refractory colorectal cancer treatment.J Transl Med. 2024 Oct 7;22(1):908. doi: 10.1186/s12967-024-05652-3. J Transl Med. 2024. PMID: 39375706 Free PMC article.
-
Role of trypanosomatid's arginase in polyamine biosynthesis and pathogenesis.Mol Biochem Parasitol. 2012 Feb;181(2):85-93. doi: 10.1016/j.molbiopara.2011.10.007. Epub 2011 Oct 20. Mol Biochem Parasitol. 2012. PMID: 22033378 Review.
Cited by
-
Druggable Metabolic Vulnerabilities Are Exposed and Masked during Progression to Castration Resistant Prostate Cancer.Biomolecules. 2022 Oct 28;12(11):1590. doi: 10.3390/biom12111590. Biomolecules. 2022. PMID: 36358940 Free PMC article. Review.
-
Myeloid-Derived Suppressor Cells (MDSC) in Melanoma Patients Treated with Anti-PD-1 Immunotherapy.Cells. 2023 Mar 2;12(5):789. doi: 10.3390/cells12050789. Cells. 2023. PMID: 36899926 Free PMC article.
-
Myeloid cells meet CD8+ T cell exhaustion in cancer: What, why and how.Chin J Cancer Res. 2024 Dec 30;36(6):616-651. doi: 10.21147/j.issn.1000-9604.2024.06.04. Chin J Cancer Res. 2024. PMID: 39802897 Free PMC article.
-
Role of amino acid metabolism in tumor immune microenvironment of colorectal cancer.Am J Cancer Res. 2025 Jan 15;15(1):233-247. doi: 10.62347/ZSOO2247. eCollection 2025. Am J Cancer Res. 2025. PMID: 39949925 Free PMC article. Review.
-
Transcending frontiers in prostate cancer: the role of oncometabolites on epigenetic regulation, CSCs, and tumor microenvironment to identify new therapeutic strategies.Cell Commun Signal. 2024 Jan 12;22(1):36. doi: 10.1186/s12964-023-01462-0. Cell Commun Signal. 2024. PMID: 38216942 Free PMC article. Review.
References
-
- Kabalin J.N., McNeal J.E., Price H.M., Freiha F.S., Stamey T.A. Unsuspected adenocarcinoma of the prostate in patients undergoing cystoprostatectomy for other causes: Incidence, histology and morphometric observations. J. Urol. 1989;141:1091–1094; discussion 1093–1094. doi: 10.1016/S0022-5347(17)41178-5. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

