Mitochondria as environments for the nuclear genome in Drosophila: mitonuclear G×G×E
- PMID: 34964900
- PMCID: PMC8851671
- DOI: 10.1093/jhered/esab066
Mitochondria as environments for the nuclear genome in Drosophila: mitonuclear G×G×E
Abstract
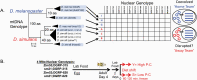
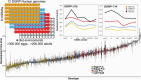
Mitochondria evolved from a union of microbial cells belonging to distinct lineages that were likely anaerobic. The evolution of eukaryotes required a massive reorganization of the 2 genomes and eventual adaptation to aerobic environments. The nutrients and oxygen that sustain eukaryotic metabolism today are processed in mitochondria through coordinated expression of 37 mitochondrial genes and over 1000 nuclear genes. This puts mitochondria at the nexus of gene-by-gene (G×G) and gene-by-environment (G×E) interactions that sustain life. Here we use a Drosophila model of mitonuclear genetic interactions to explore the notion that mitochondria are environments for the nuclear genome, and vice versa. We construct factorial combinations of mtDNA and nuclear chromosomes to test for epistatic interactions (G×G), and expose these mitonuclear genotypes to altered dietary environments to examine G×E interactions. We use development time and genome-wide RNAseq analyses to assess the relative contributions of mtDNA, nuclear chromosomes, and environmental effects on these traits (mitonuclear G×G×E). We show that the nuclear transcriptional response to alternative mitochondrial "environments" (G×G) has significant overlap with the transcriptional response of mitonuclear genotypes to altered dietary environments. These analyses point to specific transcription factors (e.g., giant) that mediated these interactions, and identified coexpressed modules of genes that may account for the overlap in differentially expressed genes. Roughly 20% of the transcriptome includes G×G genes that are concordant with G×E genes, suggesting that mitonuclear interactions are part of an organism's environment.
Keywords: coevolution; epistasis; gene expression module.
© The American Genetic Association. 2021.
Figures


Similar articles
-
Mitonuclear epistasis, genotype-by-environment interactions, and personalized genomics of complex traits in Drosophila.IUBMB Life. 2018 Dec;70(12):1275-1288. doi: 10.1002/iub.1954. Epub 2018 Nov 5. IUBMB Life. 2018. PMID: 30394643 Free PMC article.
-
Mitonuclear Interactions Mediate Transcriptional Responses to Hypoxia in Drosophila.Mol Biol Evol. 2017 Feb 1;34(2):447-466. doi: 10.1093/molbev/msw246. Mol Biol Evol. 2017. PMID: 28110272 Free PMC article.
-
Mitonuclear conflict and cooperation govern the integration of genotypes, phenotypes and environments.Philos Trans R Soc Lond B Biol Sci. 2020 Jan 20;375(1790):20190188. doi: 10.1098/rstb.2019.0188. Epub 2019 Dec 2. Philos Trans R Soc Lond B Biol Sci. 2020. PMID: 31787039 Free PMC article.
-
The role of mitonuclear incompatibilities in allopatric speciation.Cell Mol Life Sci. 2022 Jan 29;79(2):103. doi: 10.1007/s00018-021-04059-3. Cell Mol Life Sci. 2022. PMID: 35091831 Free PMC article. Review.
-
Mitonuclear interactions: evolutionary consequences over multiple biological scales.Philos Trans R Soc Lond B Biol Sci. 2014 Jul 5;369(1646):20130443. doi: 10.1098/rstb.2013.0443. Philos Trans R Soc Lond B Biol Sci. 2014. PMID: 24864313 Free PMC article. Review.
Cited by
-
Conservation Mitonuclear Replacement: Facilitated mitochondrial adaptation for a changing world.Evol Appl. 2024 Mar 10;17(3):e13642. doi: 10.1111/eva.13642. eCollection 2024 Mar. Evol Appl. 2024. PMID: 38468713 Free PMC article.
-
Mapping mitonuclear epistasis using a novel recombinant yeast population.PLoS Genet. 2023 Mar 29;19(3):e1010401. doi: 10.1371/journal.pgen.1010401. eCollection 2023 Mar. PLoS Genet. 2023. PMID: 36989278 Free PMC article.
-
Introgression and Genetic Diversity Between Two Cactophilic Drosophila (Drosophila repleta group) Species: A Case Study of an Isolated Population from the Sandstone Hills in the Southeast of Brazil.Neotrop Entomol. 2025 May 6;54(1):64. doi: 10.1007/s13744-025-01279-1. Neotrop Entomol. 2025. PMID: 40327225
-
Mitonuclear interactions shape both direct and parental effects of diet on fitness and involve a SNP in mitoribosomal 16s rRNA.PLoS Biol. 2023 Aug 21;21(8):e3002218. doi: 10.1371/journal.pbio.3002218. eCollection 2023 Aug. PLoS Biol. 2023. PMID: 37603597 Free PMC article.
-
Indirect Genetic Effects: A Cross-disciplinary Perspective on Empirical Studies.J Hered. 2022 Feb 17;113(1):1-15. doi: 10.1093/jhered/esab059. J Hered. 2022. PMID: 34643239 Free PMC article.
References
-
- Arnau-Soler A, Macdonald-Dunlop E, Adams MJ, Clarke TK, MacIntyre DJ, Milburn K, Navrady L, Hayward C, McIntosh AM, Thomson PA; Generation Scotland; Major Depressive Disorder Working Group of the Psychiatric Genomics Consortium . 2019. Genome-wide by environment interaction studies of depressive symptoms and psychosocial stress in UK Biobank and Generation Scotland. Transl Psychiatry. 9:14. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

