Single-cell RNA sequencing reveals a pro-invasive cancer-associated fibroblast subgroup associated with poor clinical outcomes in patients with gastric cancer
- PMID: 34976204
- PMCID: PMC8692898
- DOI: 10.7150/thno.60540
Single-cell RNA sequencing reveals a pro-invasive cancer-associated fibroblast subgroup associated with poor clinical outcomes in patients with gastric cancer
Abstract
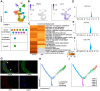
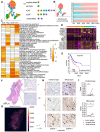
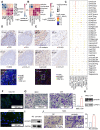
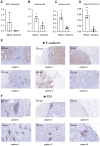
Background: The protumor activities of cancer-associated fibroblasts (CAFs) suggest that they are potential therapeutic targets for the treatment of cancer. The mechanism of CAF heterogeneity in gastric cancer (GC) remains unclear and has slowed translational advances in targeting CAFs. Therefore, a comprehensive understanding of the classification, function, activation stage, and spatial distribution of the CAF subsets in GC is urgently needed. Methods: In this study, the characteristics of the CAF subsets and the dynamic communication among the tumor microenvironment (TME) components regulated by the CAF subsets were analyzed by performing single-cell RNA sequencing of eight pairs of GC and adjacent mucosal (AM) samples. The spatial distribution of the CAF subsets in different Lauren subtypes of GC, as well as the neighborhood relations between these CAF subsets and the protumor immune cell subsets were evaluated by performing multistaining registration. Results: Tumor epithelial cells exhibited significant intratumor and intertumor variabilities, while CAFs mainly exhibited intratumor variability. Moreover, we identified four CAF subsets with different properties in GC. These four CAF subsets shared similar properties with their resident fibroblast counterparts in the adjacent mucosa but also exhibited enhanced protumor activities. Additionally, two CAF subsets, inflammatory CAFs (iCAFs) and extracellular matrix CAFs (eCAFs), communicated with adjacent immune cell subsets in the GC TME. iCAFs interacted with T cells by secreting interleukin (IL)-6 and C-X-C motif chemokine ligand 12 (CXCL12), while eCAFs correlated with M2 macrophages via the expression of periostin (POSTN). eCAFs, which function as a pro-invasive CAF subset, decreased the overall survival time of patients with GC. Conclusions: iCAFs and eCAFs not only exhibited enhanced pro-invasive activities but also mobilized the surrounding immune cells to construct a tumor-favorable microenvironment. Therefore, inhibiting their activation restrains the GC 'seed' and simultaneously improves the 'GC' soil, suggesting that it represents a promising therapeutic strategy for the treatment of GC.
Keywords: CAF heterogeneity; Gastric cancer; Multistaining registration; Tumor microenvironment; eCAF; iCAF; scRNA-Seq.
© The author(s).
Conflict of interest statement
Competing Interests: The authors have declared that no competing interest exists.
Figures



References
-
- Sung H, Ferlay J, Siegel R L, Laversanne M, Soerjomataram I, Jemal A. et al. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J Clin. 2021;71:209–49. - PubMed
-
- Smyth E C, Nilsson M, Grabsch H I, van Grieken N C, Lordick F. Gastric cancer. Lancet. 2020;396:635–48. - PubMed
-
- Lauren P. THE TWO HISTOLOGICAL MAIN TYPES OF GASTRIC CARCINOMA: DIFFUSE AND SO-CALLED INTESTINAL-TYPE CARCINOMA. AN ATTEMPT AT A HISTO-CLINICAL CLASSIFICATION. Acta Pathol Microbiol Scand. 1965;64:31–49. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

