Construction of an IS-Free Corynebacterium glutamicum ATCC 13 032 Chassis Strain and Random Mutagenesis Using the Endogenous IS Cg1 Transposase
- PMID: 34976962
- PMCID: PMC8715038
- DOI: 10.3389/fbioe.2021.751334
Construction of an IS-Free Corynebacterium glutamicum ATCC 13 032 Chassis Strain and Random Mutagenesis Using the Endogenous IS Cg1 Transposase
Abstract
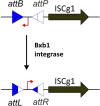
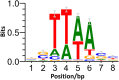
Mobile genetic elements (MGEs) contribute to instability of the host genome and plasmids. Previously, removal of the prophages in the industrial amino acid producer Corynebacterium glutamicum ATCC 13 032 resulted in strain MB001 which showed better survival under stress conditions and increased transformability. Still, eight families of Insertion Sequence (IS) elements with 27 potentially active members remain in MB001, two of which were demonstrated to be detrimental in biotechnological processes. In this study, systematical deletion of all complete IS elements in MB001 resulted in the MGE-free strain CR101. CR101 shows growth characteristics identical to the wildtype and the increased transformability of MB001. Due to its improved genome stability, we consider this strain to be an optimal host for basic research and biotechnology. As a "zero-background" host, it is also an ideal basis to study C. glutamicum IS elements. Re-sequencing of CR101 revealed that only five spontaneous point mutations had occurred during the construction process, highlighting the low mutation rate of C. glutamicum on the nucleotide level. In a second step, we developed an easily applicable ISCg1-based transposon mutagenesis system to randomly transpose a selectable marker. For optimal plasmid stability during cloning in Escherichia coli, the system utilizes a genetic switch based on the phage integrase Bxb1. Use of this integrase revealed the presence of a functional attB site in the C. glutamicum genome. To avoid cross-talk with our system and increase ease-of-use, we removed the attB site and also inserted the Bxb1 encoding gene into the chromosome of CR101. Successful insertion of single markers was verified by sequencing randomly selected mutants. Sequencing pooled mutant libraries revealed only a weak target site specificity, seemingly random distribution of insertion sites and no general strand bias. The resulting strain, ML103, together with plasmid pML10 provides a easily customizable system for random mutagenesis in an otherwise genomically stable C. glutamicum. Taken together, the MGE-free C. glutamicum strain CR101, the derivative ML103, and the plasmid pML10 provide a useful set of tools to study C. glutamicum in the future.
Keywords: IS elements; ISCg1; corynebacterium glutamcium; genetic switch; mutagenesis; prophages; “genome healing”.
Copyright © 2021 Linder , Haak , Botes , Kalinowski and Rückert .
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


Similar articles
-
Construction of a prophage-free variant of Corynebacterium glutamicum ATCC 13032 for use as a platform strain for basic research and industrial biotechnology.Appl Environ Microbiol. 2013 Oct;79(19):6006-15. doi: 10.1128/AEM.01634-13. Epub 2013 Jul 26. Appl Environ Microbiol. 2013. PMID: 23892752 Free PMC article.
-
Random mutagenesis in Corynebacterium glutamicum ATCC 13032 using an IS6100-based transposon vector identified the last unknown gene in the histidine biosynthesis pathway.BMC Genomics. 2006 Aug 10;7:205. doi: 10.1186/1471-2164-7-205. BMC Genomics. 2006. PMID: 16901339 Free PMC article.
-
Enhanced production of recombinant proteins with Corynebacterium glutamicum by deletion of insertion sequences (IS elements).Microb Cell Fact. 2015 Dec 29;14:207. doi: 10.1186/s12934-015-0401-7. Microb Cell Fact. 2015. PMID: 26715464 Free PMC article.
-
Advances and Perspectives for Genome Editing Tools of Corynebacterium glutamicum.Front Microbiol. 2021 Apr 7;12:654058. doi: 10.3389/fmicb.2021.654058. eCollection 2021. Front Microbiol. 2021. PMID: 33897668 Free PMC article. Review.
-
Plasmids in Corynebacterium glutamicum and their molecular classification by comparative genomics.J Biotechnol. 2003 Sep 4;104(1-3):27-40. doi: 10.1016/s0168-1656(03)00157-3. J Biotechnol. 2003. PMID: 12948627 Review.
Cited by
-
Comprehensive analysis of genomic variation, pan-genome and biosynthetic potential of Corynebacterium glutamicum strains.PLoS One. 2024 May 8;19(5):e0299588. doi: 10.1371/journal.pone.0299588. eCollection 2024. PLoS One. 2024. PMID: 38718091 Free PMC article.
-
Early onset of septal FtsK localization allows for efficient DNA segregation in SMC-deleted Corynebacterium glutamicum strains.mBio. 2025 Mar 12;16(3):e0285924. doi: 10.1128/mbio.02859-24. Epub 2025 Jan 28. mBio. 2025. PMID: 39873485 Free PMC article.
-
Promising non-model microbial cell factories obtained by genome reduction.Front Bioeng Biotechnol. 2024 Aug 5;12:1427248. doi: 10.3389/fbioe.2024.1427248. eCollection 2024. Front Bioeng Biotechnol. 2024. PMID: 39161352 Free PMC article. Review.
-
Bacterial genome reductions: Tools, applications, and challenges.Front Genome Ed. 2022 Aug 31;4:957289. doi: 10.3389/fgeed.2022.957289. eCollection 2022. Front Genome Ed. 2022. PMID: 36120530 Free PMC article. Review.
-
High-level recombinant protein production with Corynebacterium glutamicum using acetate as carbon source.Microb Biotechnol. 2022 Nov;15(11):2744-2757. doi: 10.1111/1751-7915.14138. Epub 2022 Sep 30. Microb Biotechnol. 2022. PMID: 36178056 Free PMC article.
References
-
- Anderson J. C. (2013). Anderson Promoter Collection. Available: http://parts.igem.org/promoters/catalog/anderson [Dataset].
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

