The exploitation of host autophagy and ubiquitin machinery by Mycobacterium tuberculosis in shaping immune responses and host defense during infection
- PMID: 35000542
- PMCID: PMC9809970
- DOI: 10.1080/15548627.2021.2021495
The exploitation of host autophagy and ubiquitin machinery by Mycobacterium tuberculosis in shaping immune responses and host defense during infection
Abstract
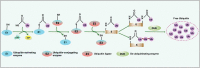
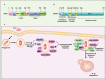
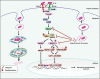
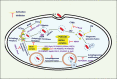
Intracellular pathogens have evolved various efficient molecular armaments to subvert innate defenses. Cellular ubiquitination, a normal physiological process to maintain homeostasis, is emerging one such exploited mechanism. Ubiquitin (Ub), a small protein modifier, is conjugated to diverse protein substrates to regulate many functions. Structurally diverse linkages of poly-Ub to target proteins allow enormous functional diversity with specificity being governed by evolutionarily conserved enzymes (E3-Ub ligases). The Ub-binding domain (UBD) and LC3-interacting region (LIR) are critical features of macroautophagy/autophagy receptors that recognize Ub-conjugated on protein substrates. Emerging evidence suggests that E3-Ub ligases unexpectedly protect against intracellular pathogens by tagging poly-Ub on their surfaces and targeting them to phagophores. Two E3-Ub ligases, PRKN and SMURF1, provide immunity against Mycobacterium tuberculosis (M. tb). Both enzymes conjugate K63 and K48-linked poly-Ub to M. tb for successful delivery to phagophores. Intriguingly, M. tb exploits virulence factors to effectively dampen host-directed autophagy utilizing diverse mechanisms. Autophagy receptors contain LIR-motifs that interact with conserved Atg8-family proteins to modulate phagophore biogenesis and fusion to the lysosome. Intracellular pathogens have evolved a vast repertoire of virulence effectors to subdue host-immunity via hijacking the host ubiquitination process. This review highlights the xenophagy-mediated clearance of M. tb involving host E3-Ub ligases and counter-strategy of autophagy inhibition by M. tb using virulence factors. The role of Ub-binding receptors and their mode of autophagy regulation is also explained. We also discuss the co-opting and utilization of the host Ub system by M. tb for its survival and virulence.Abbreviations: APC: anaphase promoting complex/cyclosome; ATG5: autophagy related 5; BCG: bacille Calmette-Guerin; C2: Ca2+-binding motif; CALCOCO2: calcium binding and coiled-coil domain 2; CUE: coupling of ubiquitin conjugation to ER degradation domains; DUB: deubiquitinating enzyme; GABARAP: GABA type A receptor-associated protein; HECT: homologous to the E6-AP carboxyl terminus; IBR: in-between-ring fingers; IFN: interferon; IL1B: interleukin 1 beta; KEAP1: kelch like ECH associated protein 1; LAMP1: lysosomal associated membrane protein 1; LGALS: galectin; LIR: LC3-interacting region; MAPK11/p38: mitogen-activated protein kinase 11; MAP1LC3/LC3: microtubule associated protein 1 light chain 3; MAP3K7/TAK1: mitogen-activated protein kinase kinase kinase 7; MAPK8/JNK: mitogen-activated protein kinase 8; MHC-II: major histocompatibility complex-II; MTOR: mechanistic target of rapamycin kinase; NBR1: NBR1 autophagy cargo receptor; NFKB1/p50: nuclear factor kappa B subunit 1; OPTN: optineurin; PB1: phox and bem 1; PE/PPE: proline-glutamic acid/proline-proline-glutamic acid; PknG: serine/threonine-protein kinase PknG; PRKN: parkin RBR E3 ubiquitin protein ligase; RBR: RING-in between RING; RING: really interesting new gene; RNF166: RING finger protein 166; ROS: reactive oxygen species; SMURF1: SMAD specific E3 ubiquitin protein ligase 1; SQSTM1: sequestosome 1; STING1: stimulator of interferon response cGAMP interactor 1; TAX1BP1: Tax1 binding protein 1; TBK1: TANK binding kinase 1; TNF: tumor necrosis factor; TRAF6: TNF receptor associated factor 6; Ub: ubiquitin; UBA: ubiquitin-associated; UBAN: ubiquitin-binding domain in ABIN proteins and NEMO; UBD: ubiquitin-binding domain; UBL: ubiquitin-like; ULK1: unc-51 like autophagy activating kinase 1.
Keywords: Autophagy; E3-Ub ligase; LC3; intracellular pathogens; phagolysosome; ubiquitin-binding receptors; virulence effectors; xenophagy.
Conflict of interest statement
No potential conflict of interest was reported by the author(s).
Figures



References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous
