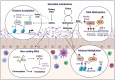
Epigenetic regulation by gut microbiota
- PMID: 35000562
- PMCID: PMC8744890
- DOI: 10.1080/19490976.2021.2022407
Epigenetic regulation by gut microbiota
Abstract
The gastrointestinal tract is continuously exposed to trillions of commensal microbes, collectively termed the microbiota, which are environmental stimuli that can direct health and disease within the host. In addition to well-established bacterial sensing pathways, microbial signals are also integrated through epigenetic modifications that calibrate the transcriptional program of host cells without altering the underlying genetic code. Microbiota-sensitive epigenetic changes include modifications to the DNA or histones, as well as regulation of non-coding RNAs. While microbiota-sensitive epigenetic mechanisms have been described in both local intestinal cells and as well in peripheral tissues, further research is required to fully decipher the complex relationship between the host and microbiota. This Review highlights current understandings of epigenetic regulation by gut microbiota and important implications of these findings in guiding therapeutic approaches to prevent or combat diseases driven by impaired microbiota-host interactions.
Keywords: HDAC; Microbiota; SCFAs; chromatin; epigenetics; epigenome; histone modification; intestine; microbiome.
Conflict of interest statement
No potential conflict of interest was reported by the author(s).
Figures
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources