First Report of bla OXA-677 with Enhanced Meropenem-Hydrolyzing Ability in Pseudomonas aeruginosa in China
- PMID: 35002263
- PMCID: PMC8725689
- DOI: 10.2147/IDR.S340662
First Report of bla OXA-677 with Enhanced Meropenem-Hydrolyzing Ability in Pseudomonas aeruginosa in China
Abstract
Purpose: OXA-10-type class D β-lactamases have shown their evolutionary potential of enhancing carbapenem resistance. This study aimed to elucidate the role of OXA-10 variants in clinical isolated multidrug resistant (MDR) Pseudomonas aeruginosa and characterize the first appearance of OXA-677 in China.
Methods: Six bla OXA-10-like-positive strains were screened by PCR from 41 P. aeruginosa strains, which were resistant to both carbapenems and ceftazidime-avibactam, collected across China in 2018. The minimum inhibitory concentrations (MIC) were determined with the broth microdilution method. The resistance-associated genes and genetic environment were investigated by whole-genome sequencing (WGS). The function and mechanism of OXA-677 β-lactamase were identified by molecular cloning and protein structure modeling.
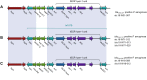
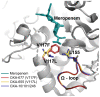
Results: All the bla OXA-10-like-positive Pseudomonas aeruginosa were MDR strains. They also had outer membrane porin defects and produced β-lactam resistance gene bla PER-1, fluoroquinolone-resistant gene crpP, aminoglycoside-resistance gene aph(3')-IIb, aph(6)-Id, aacA and aadA, fosfomycin-resistance gene fosA, sulfamethoxazole-resistance gene sul1, and chloramphenicol-resistance gene catB7. All bla OXA-10 variants were located in a Tn1403-related transposon, containing aacA4-12-bla OXA-677-aadA1, aacA4-12-bla OXA-101-aadA5, and bla OXA-246-aacA3-aadA13 gene cassette arrays, respectively. Notably, the bla OXA-677 producer showed a high MIC level of meropenem (MIC>64 mg/L). Compared to bla OXA-10, bla OXA-677 was found a G-to-T transversion at position 350, leading to a phenylalanine-for-valine substitution in position 117, which is closer to leucine155 in the omega loop of the active site. MIC of meropenem for E. coli DH5α with the recombinant plasmid pHSG398 carrying bla OXA-677 was elevated by 8 times.
Conclusion: We speculate that the OXA-10-like enzymes and the decrease of membrane permeability confer carbapenem resistance, and the V117 substitution in OXA-677 might lead to a higher resistance level of meropenem.
Keywords: CHDL; Pseudomonas aeruginosa; blaOXA-10-like gene; blaOXA-677.
© 2021 Sun et al.
Conflict of interest statement
The authors report no conflicts of interest in this work.
Figures

Similar articles
-
Characterization of acquired beta-lactamases and their genetic support in multidrug-resistant Pseudomonas aeruginosa isolates in Taiwan: the prevalence of unusual integrons.J Antimicrob Chemother. 2006 Sep;58(3):530-6. doi: 10.1093/jac/dkl266. Epub 2006 Jul 1. J Antimicrob Chemother. 2006. PMID: 16816399
-
Characteristics of rare ST463 carbapenem-resistant Pseudomonas aeruginosa clinical isolates from blood.J Glob Antimicrob Resist. 2023 Mar;32:122-130. doi: 10.1016/j.jgar.2023.01.011. Epub 2023 Feb 16. J Glob Antimicrob Resist. 2023. PMID: 36801256
-
Challenging Antimicrobial Susceptibility and Evolution of Resistance (OXA-681) during Treatment of a Long-Term Nosocomial Infection Caused by a Pseudomonas aeruginosa ST175 Clone.Antimicrob Agents Chemother. 2019 Sep 23;63(10):e01110-19. doi: 10.1128/AAC.01110-19. Print 2019 Oct. Antimicrob Agents Chemother. 2019. PMID: 31383659 Free PMC article.
-
Molecular mechanisms driving the in vivo development of OXA-10-mediated resistance to ceftolozane/tazobactam and ceftazidime/avibactam during treatment of XDR Pseudomonas aeruginosa infections.J Antimicrob Chemother. 2021 Jan 1;76(1):91-100. doi: 10.1093/jac/dkaa396. J Antimicrob Chemother. 2021. PMID: 33083833
-
Cefiderocol: A Siderophore Cephalosporin with Activity Against Carbapenem-Resistant and Multidrug-Resistant Gram-Negative Bacilli.Drugs. 2019 Feb;79(3):271-289. doi: 10.1007/s40265-019-1055-2. Drugs. 2019. PMID: 30712199 Review.
Cited by
-
Molecular mechanism of plasmid-borne resistance to sulfonamide antibiotics.Nat Commun. 2023 Jul 7;14(1):4031. doi: 10.1038/s41467-023-39778-7. Nat Commun. 2023. PMID: 37419898 Free PMC article.
-
Prioritization of Critical Factors for Surveillance of the Dissemination of Antibiotic Resistance in Pseudomonas aeruginosa: A Systematic Review.Int J Mol Sci. 2023 Oct 15;24(20):15209. doi: 10.3390/ijms242015209. Int J Mol Sci. 2023. PMID: 37894890 Free PMC article.
-
National Cohort of Compassionate Use of Meropenem-Vaborbactam: No Benefit over Meropenem for Pseudomonas aeruginosa.Antibiotics (Basel). 2024 Dec 1;13(12):1152. doi: 10.3390/antibiotics13121152. Antibiotics (Basel). 2024. PMID: 39766541 Free PMC article.
References
-
- Perez A, Gato E, Perez-Llarena J, et al. High incidence of MDR and XDR Pseudomonas aeruginosa isolates obtained from patients with ventilator-associated pneumonia in Greece, Italy and Spain as part of the MagicBullet clinical trial. J Antimicrob Chemother. 2019;74(5):1244–1252. doi:10.1093/jac/dkz030 - DOI - PubMed
-
- Yin D, Wu S, Yang Y, et al. Results from the China Antimicrobial Surveillance Network (CHINET) in 2017 of the in vitro activities of ceftazidime-avibactam and ceftolozane-tazobactam against clinical isolates of Enterobacteriaceae and Pseudomonas aeruginosa. Antimicrob Agents Chemother. 2019;63(4):e02431–18. doi:10.1128/AAC.02431-18 - DOI - PMC - PubMed
-
- Hu F, Guo Y, Zhu D, et al. CHINET surveillance of bacterial resistance: results of 2020. Chin J Infect Chemother. 2021;21(04):377–387.
LinkOut - more resources
Full Text Sources

