Environmental Tuning of Homologs of the Orange Carotenoid Protein-Encoding Gene in the Cyanobacterium Fremyella diplosiphon
- PMID: 35003049
- PMCID: PMC8739951
- DOI: 10.3389/fmicb.2021.819604
Environmental Tuning of Homologs of the Orange Carotenoid Protein-Encoding Gene in the Cyanobacterium Fremyella diplosiphon
Abstract
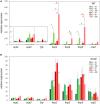
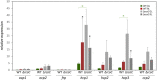
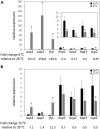
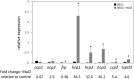
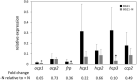
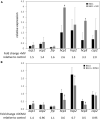
The orange carotenoid protein (OCP) family of proteins are light-activated proteins that function in dissipating excess energy absorbed by accessory light-harvesting complexes, i.e., phycobilisomes (PBSs), in cyanobacteria. Some cyanobacteria contain multiple homologs of the OCP-encoding gene (ocp). Fremyella diplosiphon, a cyanobacterium studied for light-dependent regulation of PBSs during complementary chromatic acclimation (CCA), contains several OCP homologs - two full-length OCPs, three Helical Carotenoid Proteins (HCPs) with homology to the N-terminus of OCP, and one C-terminal domain-like carotenoid protein (CCP) with homology to the C-terminus of OCP. We examined whether these homologs are distinctly regulated in response to different environmental factors, which could indicate distinct functions. We observed distinct patterns of expression for some OCP, HCP, and CCP encoding genes, and have evidence that light-dependent aspects of ocp homolog expression are regulated by photoreceptor RcaE which controls CCA. RcaE-dependent transcriptional regulator RcaC is also involved in the photoregulation of some hcp genes. Apart from light, additional environmental factors associated with cellular redox regulation impact the mRNA levels of ocp homologs, including salt, cold, and disruption of electron transport. Analyses of conserved sequences in the promoters of ocp homologs were conducted to gain additional insight into regulation of these genes. Several conserved regulatory elements were found across multiple ocp homolog promoters that potentially control differential transcriptional regulation in response to a range of environmental cues. The impact of distinct environmental cues on differential accumulation of ocp homolog transcripts indicates potential functional diversification of this gene family in cyanobacteria. These genes likely enable dynamic cellular protection in response to diverse environmental stress conditions in F. diplosiphon.
Keywords: cyanobacteria; non-photochemical quenching (NPQ); orange carotenoid protein (OCP); photosynthesis; stress.
Copyright © 2021 Petrescu, Dilbeck and Montgomery.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

References
-
- Agostoni M., Lucker B. F., Smith M. Y., Kanazawa A., Blanchard G. J., Kramer D. M., et al. (2016). Competition-based phenotyping reveals a fitness cost for maintaining phycobilisomes under fluctuating light in the cyanobacterium Fremyella diplosiphon. Algal Res 15 110–119. 10.1016/j.algal.2016.02.005 - DOI
LinkOut - more resources
Full Text Sources
Miscellaneous

