Physical Interactions With Bacteria and Protozoan Parasites Establish the Scavenger Receptor SSC4D as a Broad-Spectrum Pattern Recognition Receptor
- PMID: 35003072
- PMCID: PMC8739261
- DOI: 10.3389/fimmu.2021.760770
Physical Interactions With Bacteria and Protozoan Parasites Establish the Scavenger Receptor SSC4D as a Broad-Spectrum Pattern Recognition Receptor
Abstract
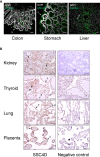
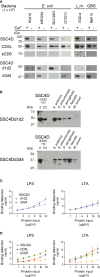
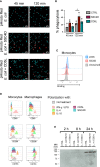
Since the pioneering discoveries, by the Nobel laureates Jules Hoffmann and Bruce Beutler, that Toll and Toll-like receptors can sense pathogenic microorganisms and initiate, in vertebrates and invertebrates, innate immune responses against microbial infections, many other families of pattern recognition receptors (PRRs) have been described. One of such receptor clusters is composed by, if not all, at least several members of the scavenger receptor cysteine-rich (SRCR) superfamily. Many SRCR proteins are plasma membrane receptors of immune cells; however, a small subset consists of secreted receptors that are therefore in circulation. We here describe the first characterization of biological and functional roles of the circulating human protein SSC4D, one of the least scrutinized members of the family. Within leukocyte populations, SSC4D was found to be expressed by monocytes/macrophages, neutrophils, and B cells, but its production was particularly evident in epithelial cells of several organs and tissues, namely, in the kidney, thyroid, lung, placenta, intestinal tract, and liver. Similar to other SRCR proteins, SSC4D shows the capacity of physically binding to different species of bacteria, and this opsonization can increase the phagocytic capacity of monocytes. Importantly, we have uncovered the capacity of SSC4D of binding to several protozoan parasites, a singular feature seldom described for PRRs in general and here demonstrated for the first time for an SRCR family member. Overall, our study is pioneer in assigning a PRR role to SSC4D.
Keywords: bacteria; circulating receptors; parasites; pattern recognition receptors; scavenger receptor cysteine-rich.
Copyright © 2021 Cardoso, Santos, Almeida, Sá, Pérez-Cabezas, Oliveira, Tavares and Carmo.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

