Single-cell RNA Seq reveals cellular landscape-specific characteristics and potential etiologies for adolescent idiopathic scoliosis
- PMID: 35005449
- PMCID: PMC8717101
- DOI: 10.1002/jsp2.1184
Single-cell RNA Seq reveals cellular landscape-specific characteristics and potential etiologies for adolescent idiopathic scoliosis
Abstract
Backgrounds: Abnormal vertebral growth and development have been found in adolescent idiopathic scoliosis (AIS) patients, and the proliferation and differentiation of bone development-related cells play important roles in its pathogenesis. However, a comprehensive single-cell-level differentiation roadmap in AIS has not been achieved.
Methods: The present study compared the single-cell level cellular landscapes of spinal cancellous bone tissues between AIS patients and healthy subjects using high throughput single-cell RNA sequencing (scRNA-seq), which covers multiple cellular lineages including osteoblast, chondrocyte, osteoclast and related immunocytes. We constructed the differentiation trajectories of bone development-related cell lineages through pseudotime analysis, and the intercellular-communication networks between bone development-related cells and immunocytes were further developed.
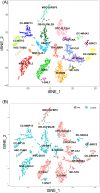
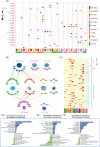
Results: A total of 11 distinct cell clusters were identified according to the genome-wide transcriptome profiles. t-Distributed stochastic neighbor embedding (t-SNE) analysis showed that mesenchymal stem cells (MSC) were classified into three subtypes: MSC-LOXL2, MSC-IGFBP5, and MSC-GJA1. Gene ontology (GO) analysis showed that MSC-GJA1 might possess greater osteoblast differentiation potential than the others. MSC-IGFBP5 was the specific MSC subtype observed only in AIS. There were two distinct gene expression clusters: OB-DPT and OB-OLFML2B, and the counts of osteoblasts derived from AIS was significantly less than that of non-AIS subjects. In AIS patients, MSC-IGFBP5 failed to differentiate into osteoblasts and exhibited negative regulation of cell proliferation and enhanced cell death. CPC-PCNA was found to be the specific chondrocyte progenitor cell (CPC) subtype observed only in AIS patients. The cell counts of OC-BIRC3 in AIS were less than those in controls. Pseudotime analysis suggested two possible distinct osteoclast differentiation patterns in AIS and control subjects. Monocytes in AIS mainly differentiated into OC-CRISP3.
Conclusions: Our single-cell analysis first revealed differences existed in the cellular states between AIS patients and healthy subjects and found the differentiation disruption of specific MSC and CPC clusters in AIS. Cell communication analysis provided the possible pathogenesis of osteoblast and chondrocyte differentiation dysfunction in AIS.
Keywords: adolescent idiopathic scoliosis; differentiation; etiology; single‐cell RNA sequencing.
© 2021 The Authors. JOR Spine published by Wiley Periodicals LLC on behalf of Orthopaedic Research Society.
Figures






References
-
- Grossman DC, Curry SJ, Owens DK, et al. Screening for adolescent idiopathic scoliosis: US preventive services task force recommendation statement. JAMA. 2018;319(2):165‐172. - PubMed
-
- Kouwenhoven JW, Castelein RM. The pathogenesis of adolescent idiopathic scoliosis: review of the literature. Spine. 2008;33(26):2898‐2908. - PubMed
LinkOut - more resources
Full Text Sources
Miscellaneous

