Biodistribution of unmodified cardiosphere-derived cell extracellular vesicles using single RNA tracing
- PMID: 35005847
- PMCID: PMC8743874
- DOI: 10.1002/jev2.12178
Biodistribution of unmodified cardiosphere-derived cell extracellular vesicles using single RNA tracing
Abstract
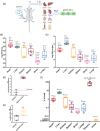
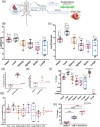
Extracellular vesicles (EVs) are potent signalling mediators. Although interest in EV translation is ever-increasing, development efforts are hampered by the inability to reliably assess the uptake of EVs and their RNA cargo. Here, we establish a novel qPCR-based method for the detection of unmodified EVS using an RNA Tracer (DUST). In this proof-of-concept study we use a human-specific Y RNA-derived small RNA (YsRNA) we dub "NT4" that is enriched in cardiosphere-derived cell small EVs (CDC-sEVs). The assay is robust, sensitive, and reproducible. Intravenously administered CDC-sEVs accumulated primarily in the heart on a per mg basis. Cardiac injury enhanced EV uptake in the heart, liver, and brain. Inhibition of EV docking by heparin suppressed uptake variably, while inhibition of endocytosis attenuated uptake in all organs. In vitro, EVs were uptaken more efficiently by macrophages, endothelial cells, and cardiac fibroblasts compared to cardiomyocytes. These findings demonstrate the utility of DUST to assess uptake of EVs in vivo and in vitro.
Keywords: Y-derived small RNA cardiosphere-derived cells; YsRNA; biodistribution; extracellular vesicles; qPCR; small non-coding RNA.
© 2021 The Authors. Journal of Extracellular Vesicles published by Wiley Periodicals, LLC on behalf of the International Society for Extracellular Vesicles.
Figures




Similar articles
-
Mononuclear phagocyte system blockade using extracellular vesicles modified with CD47 on membrane surface for myocardial infarction reperfusion injury treatment.Biomaterials. 2021 Aug;275:121000. doi: 10.1016/j.biomaterials.2021.121000. Epub 2021 Jun 28. Biomaterials. 2021. PMID: 34218049
-
Y RNA fragment in extracellular vesicles confers cardioprotection via modulation of IL-10 expression and secretion.EMBO Mol Med. 2017 Mar;9(3):337-352. doi: 10.15252/emmm.201606924. EMBO Mol Med. 2017. PMID: 28167565 Free PMC article.
-
Mechanistic and therapeutic distinctions between cardiosphere-derived cell and mesenchymal stem cell extracellular vesicle non-coding RNA.Sci Rep. 2021 Apr 21;11(1):8666. doi: 10.1038/s41598-021-87939-9. Sci Rep. 2021. PMID: 33883598 Free PMC article.
-
Cardiac Extracellular Vesicles in Normal and Infarcted Heart.Int J Mol Sci. 2016 Jan 5;17(1):63. doi: 10.3390/ijms17010063. Int J Mol Sci. 2016. PMID: 26742038 Free PMC article. Review.
-
Extracellular Vesicles as Mediators of Cellular Crosstalk Between Immune System and Kidney Graft.Front Immunol. 2020 Feb 27;11:74. doi: 10.3389/fimmu.2020.00074. eCollection 2020. Front Immunol. 2020. PMID: 32180768 Free PMC article. Review.
Cited by
-
Tracking tools of extracellular vesicles for biomedical research.Front Bioeng Biotechnol. 2022 Nov 18;10:943712. doi: 10.3389/fbioe.2022.943712. eCollection 2022. Front Bioeng Biotechnol. 2022. PMID: 36466335 Free PMC article. Review.
-
Extracellular Non-Coding RNAs in Cardiovascular Diseases.Pharmaceutics. 2023 Jan 3;15(1):155. doi: 10.3390/pharmaceutics15010155. Pharmaceutics. 2023. PMID: 36678784 Free PMC article. Review.
-
Minimal information for studies of extracellular vesicles (MISEV2023): From basic to advanced approaches.J Extracell Vesicles. 2024 Feb;13(2):e12404. doi: 10.1002/jev2.12404. J Extracell Vesicles. 2024. PMID: 38326288 Free PMC article.
-
TDO2-augmented fibroblasts secrete EVs enriched in immunomodulatory Y-derived small RNA.J Extracell Biol. 2023 Feb 2;2(2):e73. doi: 10.1002/jex2.73. eCollection 2023 Feb. J Extracell Biol. 2023. PMID: 38938522 Free PMC article.
-
Oral Bioavailability of a Noncoding RNA Drug, TY1, That Acts on Macrophages.J Extracell Biol. 2025 Aug 14;4(8):e70081. doi: 10.1002/jex2.70081. eCollection 2025 Aug. J Extracell Biol. 2025. PMID: 40823342 Free PMC article.
References
-
- Bruno, V. , Svensson‐Arvelund, J. , Rubér, M. , Berg, G. , Piccione, E. , Jenmalm, M. C. , & Ernerudh, J. (2018).Effects of low molecular weight heparin on the polarization and cytokine profile of macrophages and T helper cells in vitro. Scientific Reports 8, 4166, 10.1038/s41598-018-22418-2 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

