Structural Insights into Dihydroxylation of Terephthalate, a Product of Polyethylene Terephthalate Degradation
- PMID: 35007143
- PMCID: PMC8923216
- DOI: 10.1128/JB.00543-21
Structural Insights into Dihydroxylation of Terephthalate, a Product of Polyethylene Terephthalate Degradation
Abstract
Biodegradation of terephthalate (TPA) is a highly desired catabolic process for the bacterial utilization of this polyethylene terephthalate (PET) depolymerization product, but to date, the structure of terephthalate dioxygenase (TPDO), a Rieske oxygenase (RO) that catalyzes the dihydroxylation of TPA to a cis-diol, is unavailable. In this study, we characterized the steady-state kinetics and first crystal structure of TPDO from Comamonas testosteroni KF1 (TPDOKF1). TPDOKF1 exhibited substrate specificity for TPA (kcat/Km = 57 ± 9 mM-1 s-1). The TPDOKF1 structure harbors characteristic RO features as well as a unique catalytic domain that rationalizes the enzyme's function. The docking and mutagenesis studies reveal that its substrate specificity for TPA is mediated by the Arg309 and Arg390 residues, positioned on opposite faces of the active site. Additionally, residue Gln300 is also proven to be crucial for the activity, as its mutation to alanine decreases the activity (kcat) by 80%. This study delineates the structural features that dictate the substrate recognition and specificity of TPDO. IMPORTANCE Global plastic pollution has become the most pressing environmental issue. Recent studies on enzymes depolymerizing polyethylene terephthalate plastic into terephthalate (TPA) show some potential for tackling this. Microbial utilization of this released product, TPA, is an emerging and promising strategy for waste-to-value creation. Research in the last decade has identified terephthalate dioxygenase (TPDO) as being responsible for initiating the enzymatic degradation of TPA in a few Gram-negative and Gram-positive bacteria. Here, we determined the crystal structure of TPDO from Comamonas testosteroni KF1 and revealed that it possesses a unique catalytic domain featuring two basic residues in the active site to recognize TPA. Biochemical and mutagenesis studies demonstrated the crucial residues responsible for the substrate specificity of this enzyme.
Keywords: Comamonas testosteroni KF1; Rieske center; mononuclear iron; polyethylene terephthalate; terephthalate dioxygenase.
Conflict of interest statement
The authors declare no conflict of interest.
Figures





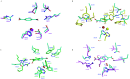
Similar articles
-
Molecular insights into substrate recognition and catalysis by phthalate dioxygenase from Comamonas testosteroni.J Biol Chem. 2021 Dec;297(6):101416. doi: 10.1016/j.jbc.2021.101416. Epub 2021 Nov 17. J Biol Chem. 2021. PMID: 34800435 Free PMC article.
-
[Advances in the structure and function of MHETase].Sheng Wu Gong Cheng Xue Bao. 2024 Sep 25;40(9):2812-2830. doi: 10.13345/j.cjb.230791. Sheng Wu Gong Cheng Xue Bao. 2024. PMID: 39319709 Review. Chinese.
-
Expression, purification, kinetics, and crystallization of non-heme mononuclear iron enzymes: Biphenyl, Phthalate, and Terephthalate dioxygenases.Methods Enzymol. 2024;704:39-58. doi: 10.1016/bs.mie.2024.05.014. Epub 2024 Jun 8. Methods Enzymol. 2024. PMID: 39300656
-
Characterization of the terephthalate degradation genes of Comamonas sp. strain E6.Appl Environ Microbiol. 2006 Mar;72(3):1825-32. doi: 10.1128/AEM.72.3.1825-1832.2006. Appl Environ Microbiol. 2006. PMID: 16517628 Free PMC article.
-
Microbial degradation and valorization of poly(ethylene terephthalate) (PET) monomers.World J Microbiol Biotechnol. 2022 Apr 15;38(5):89. doi: 10.1007/s11274-022-03270-z. World J Microbiol Biotechnol. 2022. PMID: 35426614 Review.
Cited by
-
Functional and spectroscopic approaches to determining thermal limitations of Rieske oxygenases.Methods Enzymol. 2024;703:299-328. doi: 10.1016/bs.mie.2024.05.021. Epub 2024 Jun 29. Methods Enzymol. 2024. PMID: 39261001 Free PMC article.
-
Custom tuning of Rieske oxygenase reactivity.Nat Commun. 2023 Sep 20;14(1):5858. doi: 10.1038/s41467-023-41428-x. Nat Commun. 2023. PMID: 37730711 Free PMC article.
-
Leveraging a Structural Blueprint to Rationally Engineer the Rieske Oxygenase TsaM.Biochemistry. 2023 Jun 6;62(11):1807-1822. doi: 10.1021/acs.biochem.3c00150. Epub 2023 May 15. Biochemistry. 2023. PMID: 37188334 Free PMC article.
-
Engineering Rieske oxygenase activity one piece at a time.Curr Opin Chem Biol. 2023 Feb;72:102227. doi: 10.1016/j.cbpa.2022.102227. Epub 2022 Nov 18. Curr Opin Chem Biol. 2023. PMID: 36410250 Free PMC article. Review.
-
Contrasting Mechanisms of Aromatic and Aryl-Methyl Substituent Hydroxylation by the Rieske Monooxygenase Salicylate 5-Hydroxylase.Biochemistry. 2023 Jan 17;62(2):507-523. doi: 10.1021/acs.biochem.2c00610. Epub 2022 Dec 30. Biochemistry. 2023. PMID: 36583545 Free PMC article.
References
-
- Knott BC, Erickson E, Allen MD, Gado JE, Graham R, Kearns FL, Pardo I, Topuzlu E, Anderson JJ, Austin HP, Dominick G, Johnson CW, Rorrer NA, Szostkiewicz CJ, Copié V, Payne CM, Woodcock HL, Donohoe BS, Beckham GT, McGeehan JE. 2020. Characterization and engineering of a two-enzyme system for plastics depolymerization. Proc Natl Acad Sci USA 117:25476–25485. 10.1073/pnas.2006753117. - DOI - PMC - PubMed
-
- Austin HP, Allen MD, Donohoe BS, Rorrer NA, Kearns FL, Silveira RL, Pollard BC, Dominick G, Duman R, El Omari K, Mykhaylyk V, Wagner A, Michener WE, Amore A, Skaf MS, Crowley MF, Thorne AW, Johnson CW, Woodcock HL, McGeehan JE, Beckham GT. 2018. Characterization and engineering of a plastic-degrading aromatic polyesterase. Proc Natl Acad Sci USA 115:E4350–E4357. 10.1073/pnas.1718804115. - DOI - PMC - PubMed
-
- Pardo I, Jha RK, Bermel RE, Bratti F, Gaddis M, McIntyre E, Michener W, Neidle EL, Dale T, Beckham GT, Johnson CW. 2020. Gene amplification, laboratory evolution, and biosensor screening reveal MucK as a terephthalic acid transporter in Acinetobacter baylyi ADP1. Metab Eng 62:260–274. 10.1016/j.ymben.2020.09.009. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

