Downregulation of Methionine Cycle Genes MAT1A and GNMT Enriches Protein-Associated Translation Process and Worsens Hepatocellular Carcinoma Prognosis
- PMID: 35008908
- PMCID: PMC8745498
- DOI: 10.3390/ijms23010481
Downregulation of Methionine Cycle Genes MAT1A and GNMT Enriches Protein-Associated Translation Process and Worsens Hepatocellular Carcinoma Prognosis
Abstract
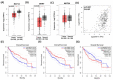
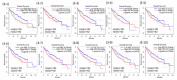
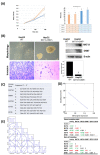
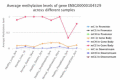
The major biological methyl donor, S-adenosylmethionine (adoMet) synthesis occurs mainly in the liver. Methionine adenosyltransferase 1A (MAT1A) and glycine N-methyltransferase (GNMT) are two key enzymes involved in the functional implications of that variation. We collected 42 RNA-seq data from paired hepatocellular carcinoma (HCC) and its adjacent normal liver tissue from the Cancer Genome Atlas (TCGA). There was no mutation found in MAT1A or GNMT RNA in the 42 HCC patients. The 11,799 genes were annotated in the RNA-Seq data, and their expression levels were used to investigate the phenotypes of low MAT1A and low GNMT by Gene Set Enrichment Analysis (GSEA). The REACTOME_TRANSLATION gene set was enriched and visualized in a heatmap along with corresponding differences in gene expression between low MAT1A versus high MAT1A and low GNMT versus high GNMT. We identified 43 genes of the REACTOME_TRANSLATION gene set that are powerful prognosis factors in HCC. The significantly predicted genes were referred into eukaryotic translation initiation (EIF3B, EIF3K), eukaryotic translation elongation (EEF1D), and ribosomal proteins (RPs). Cell models expressing various MAT1A and GNMT proved that simultaneous restoring the expression of MAT1A and GNMT decreased cell proliferation, invasion, as well as the REACTOME_TRANSLATION gene EEF1D, consistent with a better prognosis in human HCC. We demonstrated new findings that downregulation or defect in MAT1A and GNMT genes can enrich the protein-associated translation process that may account for poor HCC prognosis. This is the first study demonstrated that MAT1A and GNMT, the 2 key enzymes involved in methionine cycle, could attenuate the function of ribosome translation. We propose a potential novel mechanism by which the diminished GNMT and MAT1A expression may confer poor prognosis for HCC.
Keywords: GNMT; MAT1A; human hepatocellular carcinoma; methionine cycle.
Conflict of interest statement
There is no conflict of interest in all authors.
Figures



Similar articles
-
Human liver methionine cycle: MAT1A and GNMT gene resequencing, functional genomics, and hepatic genotype-phenotype correlation.Drug Metab Dispos. 2012 Oct;40(10):1984-92. doi: 10.1124/dmd.112.046953. Epub 2012 Jul 17. Drug Metab Dispos. 2012. PMID: 22807109 Free PMC article.
-
Pleiotropic effects of methionine adenosyltransferases deregulation as determinants of liver cancer progression and prognosis.J Hepatol. 2013 Oct;59(4):830-41. doi: 10.1016/j.jhep.2013.04.031. Epub 2013 May 7. J Hepatol. 2013. PMID: 23665184 Review.
-
Role of transcriptional and posttranscriptional regulation of methionine adenosyltransferases in liver cancer progression.Hepatology. 2012 Jul;56(1):165-75. doi: 10.1002/hep.25643. Epub 2012 Jun 5. Hepatology. 2012. PMID: 22318685
-
Glycine-N methyltransferase expression in HepG2 cells is involved in methyl group homeostasis by regulating transmethylation kinetics and DNA methylation.J Nutr. 2011 May;141(5):777-82. doi: 10.3945/jn.110.135954. Epub 2011 Mar 16. J Nutr. 2011. PMID: 21411609
-
S-adenosylmethionine in liver health, injury, and cancer.Physiol Rev. 2012 Oct;92(4):1515-42. doi: 10.1152/physrev.00047.2011. Physiol Rev. 2012. PMID: 23073625 Free PMC article. Review.
Cited by
-
The oncogenic mechanisms of the Janus kinase-signal transducer and activator of transcription pathway in digestive tract tumors.Cell Commun Signal. 2024 Jan 25;22(1):68. doi: 10.1186/s12964-023-01421-9. Cell Commun Signal. 2024. PMID: 38273295 Free PMC article. Review.
-
Exploration of Potential Biomarkers and Immune Landscape for Hepatoblastoma: Evidence from Machine Learning Algorithm.Evid Based Complement Alternat Med. 2022 Jul 31;2022:2417134. doi: 10.1155/2022/2417134. eCollection 2022. Evid Based Complement Alternat Med. 2022. Retraction in: Evid Based Complement Alternat Med. 2023 Oct 18;2023:9893765. doi: 10.1155/2023/9893765. PMID: 35958911 Free PMC article. Retracted.
-
Translocation of Methionine Adenosyl Transferase MAT2A and Its Prognostic Relevance for Liver Hepatocellular Carcinoma.Int J Mol Sci. 2023 May 22;24(10):9103. doi: 10.3390/ijms24109103. Int J Mol Sci. 2023. PMID: 37240447 Free PMC article.
-
Association of metallothionein 2A rs10636 with low mean corpuscular volume (MCV), low mean corpuscular haemoglobin (MCH) in healthy Taiwanese.Sci Rep. 2023 Jan 23;13(1):1292. doi: 10.1038/s41598-022-27304-6. Sci Rep. 2023. PMID: 36690679 Free PMC article.
-
Ketogenic Diet Consumption Inhibited Mitochondrial One-Carbon Metabolism.Int J Mol Sci. 2022 Mar 26;23(7):3650. doi: 10.3390/ijms23073650. Int J Mol Sci. 2022. PMID: 35409009 Free PMC article.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

