A Novel Hyperactive Nud1 Mitotic Exit Network Scaffold Causes Spindle Position Checkpoint Bypass in Budding Yeast
- PMID: 35011608
- PMCID: PMC8750578
- DOI: 10.3390/cells11010046
A Novel Hyperactive Nud1 Mitotic Exit Network Scaffold Causes Spindle Position Checkpoint Bypass in Budding Yeast
Abstract
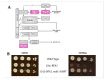
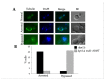
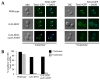
Mitotic exit is a critical cell cycle transition that requires the careful coordination of nuclear positioning and cyclin B destruction in budding yeast for the maintenance of genome integrity. The mitotic exit network (MEN) is a Ras-like signal transduction pathway that promotes this process during anaphase. A crucial step in MEN activation occurs when the Dbf2-Mob1 protein kinase complex associates with the Nud1 scaffold protein at the yeast spindle pole bodies (SPBs; centrosome equivalents) and thereby becomes activated. This requires prior priming phosphorylation of Nud1 by Cdc15 at SPBs. Cdc15 activation, in turn, requires both the Tem1 GTPase and the Polo kinase Cdc5, but how Cdc15 associates with SPBs is not well understood. We have identified a hyperactive allele of NUD1, nud1-A308T, that recruits Cdc15 to SPBs in all stages of the cell cycle in a CDC5-independent manner. This allele leads to early recruitment of Dbf2-Mob1 during metaphase and requires known Cdc15 phospho-sites on Nud1. The presence of nud1-A308T leads to loss of coupling between nuclear position and mitotic exit in cells with mispositioned spindles. Our findings highlight the importance of scaffold regulation in signaling pathways to prevent improper activation.
Keywords: Cdc15; Dbf2; MEN; Mob1; Nud1; mitotic exit; spindle position checkpoint.
Conflict of interest statement
The authors declare no conflict of interest.
Figures







References
-
- Sudakin V., Ganoth D., Dahan A., Heller H., Hershko J., Luca F.C., Ruderman J.V., Hershko A. The cyclosome, a large complex containing cyclin-selective ubiquitin ligase activity, targets cyclins for destruction at the end of mitosis. Mol. Biol. Cell. 1995;6:185–197. doi: 10.1091/mbc.6.2.185. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

