Comprehensive Molecular Landscape of Cetuximab Resistance in Head and Neck Cancer Cell Lines
- PMID: 35011716
- PMCID: PMC8750399
- DOI: 10.3390/cells11010154
Comprehensive Molecular Landscape of Cetuximab Resistance in Head and Neck Cancer Cell Lines
Abstract
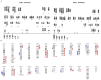
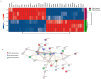
Cetuximab is the sole anti-EGFR monoclonal antibody that is FDA approved to treat head and neck squamous cell carcinoma (HNSCC). However, no predictive biomarkers of cetuximab response are known for HNSCC. Herein, we address the molecular mechanisms underlying cetuximab resistance in an in vitro model. We established a cetuximab resistant model (FaDu), using increased cetuximab concentrations for more than eight months. The resistance and parental cells were evaluated for cell viability and functional assays. Protein expression was analyzed by Western blot and human cell surface panel by lyoplate. The mutational profile and copy number alterations (CNA) were analyzed using whole-exome sequencing (WES) and the NanoString platform. FaDu resistant clones exhibited at least two-fold higher IC50 compared to the parental cell line. WES showed relevant mutations in several cancer-related genes, and the comparative mRNA expression analysis showed 36 differentially expressed genes associated with EGFR tyrosine kinase inhibitors resistance, RAS, MAPK, and mTOR signaling. Importantly, we observed that overexpression of KRAS, RhoA, and CD44 was associated with cetuximab resistance. Protein analysis revealed EGFR phosphorylation inhibition and mTOR increase in resistant cells. Moreover, the resistant cell line demonstrated an aggressive phenotype with a significant increase in adhesion, the number of colonies, and migration rates. Overall, we identified several molecular alterations in the cetuximab resistant cell line that may constitute novel biomarkers of cetuximab response such as mTOR and RhoA overexpression. These findings indicate new strategies to overcome anti-EGFR resistance in HNSCC.
Keywords: EGFR; biomarkers; cetuximab; drug resistance; head and neck tumors; in vitro; pre-clinical.
Conflict of interest statement
The authors declare no conflict of interest.
Figures





References
-
- de Carvalho A.C., Perdomo S., dos Santos W., Fernandes G.C., de Jesus L.M., Carvalho R.S., Scapulatempo-Neto C., de Almeida G.C., Sorroche B.P., Arantes L.M.R.B., et al. Impact of genetic variants in clinical outcome of a cohort of patients with oropharyngeal squamous cell carcinoma. Sci. Rep. 2020;10:1–12. doi: 10.1038/s41598-020-66741-z. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

