Lipidomic profiling of human serum enables detection of pancreatic cancer
- PMID: 35013261
- PMCID: PMC8748654
- DOI: 10.1038/s41467-021-27765-9
Lipidomic profiling of human serum enables detection of pancreatic cancer
Abstract
Pancreatic cancer has the worst prognosis among all cancers. Cancer screening of body fluids may improve the survival time prognosis of patients, who are often diagnosed too late at an incurable stage. Several studies report the dysregulation of lipid metabolism in tumor cells, suggesting that changes in the blood lipidome may accompany tumor growth. Here we show that the comprehensive mass spectrometric determination of a wide range of serum lipids reveals statistically significant differences between pancreatic cancer patients and healthy controls, as visualized by multivariate data analysis. Three phases of biomarker discovery research (discovery, qualification, and verification) are applied for 830 samples in total, which shows the dysregulation of some very long chain sphingomyelins, ceramides, and (lyso)phosphatidylcholines. The sensitivity and specificity to diagnose pancreatic cancer are over 90%, which outperforms CA 19-9, especially at an early stage, and is comparable to established diagnostic imaging methods. Furthermore, selected lipid species indicate a potential as prognostic biomarkers.
© 2022. The Author(s).
Conflict of interest statement
M.H., E.C., R.J., and D.W. are listed as inventors on patent EP 3514545 related to this work. All other authors declare no competing interests.
Figures



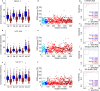





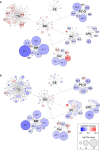
References
-
- Ryan DP, Hong TS, Bardeesy N. Pancreatic adenocarcinoma. N. Engl. J. Med. 2014;371:1039–1049. - PubMed
-
- American cancer society. Cancer treatment & survivorship facts & figures. https://www.cancer.org/research/cancer-facts-statistics/survivor-facts-f... (2016–2017).
-
- Toft J, et al. Imaging modalities in the diagnosis of pancreatic adenocarcinoma: A systematic review and meta-analysis of sensitivity, specificity, and diagnostic accuracy. Eur. J. Radiol. 2017;92:17–23. - PubMed
-
- Duffy MJ, et al. Tumor markers in pancreatic cancer: a European group on tumor markers (EGTM) status report. Ann. Oncol. 2010;21:441–447. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

