Systematic Variation of Both the Aromatic Cage and Dialkyllysine via GCE-SAR Reveal Mechanistic Insights in CBX5 Reader Protein Binding
- PMID: 35014255
- PMCID: PMC9048841
- DOI: 10.1021/acs.jmedchem.1c02049
Systematic Variation of Both the Aromatic Cage and Dialkyllysine via GCE-SAR Reveal Mechanistic Insights in CBX5 Reader Protein Binding
Abstract
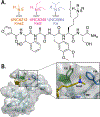
Development of inhibitors for histone methyllysine reader proteins is an active area of research due to the importance of reader protein-methyllysine interactions in transcriptional regulation and disease. Optimized peptide-based chemical probes targeting methyllysine readers favor larger alkyllysine residues in place of methyllysine. However, the mechanism by which these larger substituents drive tighter binding is not well understood. This study describes the development of a two-pronged approach combining genetic code expansion (GCE) and structure-activity relationships (SAR) through systematic variation of both the aromatic binding pocket in the protein and the alkyllysine residues in the peptide to probe inhibitor recognition in the CBX5 chromodomain. We demonstrate a novel change in driving force for larger alkyllysines, which weaken cation-π interactions but increases dispersion forces, resulting in tighter binding. This GCE-SAR approach establishes discrete energetic contributions to binding from both ligand and protein, providing a powerful tool to gain mechanistic understanding of SAR trends.
Conflict of interest statement
The authors declare no competing financial interest.
Figures

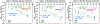



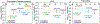
References
-
- Dawson MA; Kouzarides T; Huntly BJ, Targeting epigenetic readers in cancer. N. Engl. J. Med. 2012, 367 (7), 647–657. - PubMed
-
- Helin K; Dhanak D, Chromatin proteins and modifications as drug targets. Nature 2013, 502 (7472), 480–488. - PubMed
-
- McGrath J; Trojer P, Targeting histone lysine methylation in cancer. Pharmacol. Ther. 2015, 150, 1–22. - PubMed
-
- Murn J; Shi Y, The winding path of protein methylation research: milestones and new frontiers. Nat. Rev. Mol. Cell Biol. 2017, 18 (8), 517–527. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

