2-Hydroxyglutarate destabilizes chromatin regulatory landscape and lineage fidelity to promote cellular heterogeneity
- PMID: 35021081
- PMCID: PMC8811753
- DOI: 10.1016/j.celrep.2021.110220
2-Hydroxyglutarate destabilizes chromatin regulatory landscape and lineage fidelity to promote cellular heterogeneity
Abstract
The epigenome delineates lineage-specific transcriptional programs and restricts cell plasticity to prevent non-physiological cell fate transitions. Although cell diversification fosters tumor evolution and therapy resistance, upstream mechanisms that regulate the stability and plasticity of the cancer epigenome remain elusive. Here we show that 2-hydroxyglutarate (2HG) not only suppresses DNA repair but also mediates the high-plasticity chromatin landscape. A combination of single-cell epigenomics and multi-omics approaches demonstrates that 2HG disarranges otherwise well-preserved stable nucleosome positioning and promotes cell-to-cell variability. 2HG induces loss of motif accessibility to the luminal-defining transcriptional factors FOXA1, FOXP1, and GATA3 and a shift from luminal to basal-like gene expression. Breast tumors with high 2HG exhibit enhanced heterogeneity with undifferentiated epigenomic signatures linked to adverse prognosis. Further, ascorbate-2-phosphate (A2P) eradicates heterogeneity and impairs growth of high 2HG-producing breast cancer cells. These findings suggest 2HG as a key determinant of cancer plasticity and provide a rational strategy to counteract tumor cell evolution.
Keywords: 2-hydroxyglutarate; DNA repair; breast cancer; cancer cell plasticity; chromatin CyTOF; epigenome fluctuation; lineage fidelity; luminal-to-basal transition; oncometabolite; single-cell epigenomics.
Published by Elsevier Inc.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures


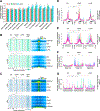




References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

