Lysoptosis is an evolutionarily conserved cell death pathway moderated by intracellular serpins
- PMID: 35022507
- PMCID: PMC8755814
- DOI: 10.1038/s42003-021-02953-x
Lysoptosis is an evolutionarily conserved cell death pathway moderated by intracellular serpins
Abstract
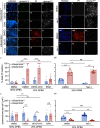
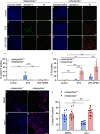
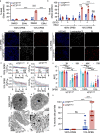
Lysosomal membrane permeabilization (LMP) and cathepsin release typifies lysosome-dependent cell death (LDCD). However, LMP occurs in most regulated cell death programs suggesting LDCD is not an independent cell death pathway, but is conscripted to facilitate the final cellular demise by other cell death routines. Previously, we demonstrated that Caenorhabditis elegans (C. elegans) null for a cysteine protease inhibitor, srp-6, undergo a specific LDCD pathway characterized by LMP and cathepsin-dependent cytoplasmic proteolysis. We designated this cell death routine, lysoptosis, to distinguish it from other pathways employing LMP. In this study, mouse and human epithelial cells lacking srp-6 homologues, mSerpinb3a and SERPINB3, respectively, demonstrated a lysoptosis phenotype distinct from other cell death pathways. Like in C. elegans, this pathway depended on LMP and released cathepsins, predominantly cathepsin L. These studies suggested that lysoptosis is an evolutionarily-conserved eukaryotic LDCD that predominates in the absence of neutralizing endogenous inhibitors.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures







References
-
- Kreuzaler PA, et al. Stat3 controls lysosomal-mediated cell death in vivo. Nat. Cell Biol. 2011;13:303–309. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

