Characterization of Cyclophilin from Thaumarchaeota Nitrosopumilus maritimus: Implications on the Diversity of Chaperone-like Activity in the Archaeal Domain
- PMID: 35036680
- PMCID: PMC8756454
- DOI: 10.1021/acsomega.1c03216
Characterization of Cyclophilin from Thaumarchaeota Nitrosopumilus maritimus: Implications on the Diversity of Chaperone-like Activity in the Archaeal Domain
Abstract
The Archaea constitute separate domain of life and show resemblance with bacteria in their metabolic pathways while showing similarity with eukaryotes at the level of molecular processes such as cell division, DNA replication, protein synthesis, and proteostasis. However, the molecular machinery of archaea can be considered a simpler version of that found in eukaryotes because of the absence of multiple paralogs for any given molecular factor. Therefore, archaeal systems can possibly be used as a model system for understanding the eukaryotic protein folding machinery and thereby may help to address the molecular mechanism of various protein (mis)foldings and diseases. In the process of protein folding, the cis-trans isomerization of the peptide-prolyl bond is a rate-limiting step for the correct folding of proteins. Different types of peptidyl-prolyl cis-trans isomerases can accelerate this reaction, e.g., cyclophilin, FKBP, and parvulin. Among the five phyla of the archaeal domain, homologs of the cyclophilin protein are found only in two. Here we have characterized a cyclophilin from an archaeal organism, Nitrosopumilus maritimus (NmCyp), belonging to the phylum Thaumarchaeota. Like other known cyclophilins, NmCyp also possesses PPIase activity that can be inhibited by cyclosporine A. Generally, archaeal proteins are expected to possess differential thermal stability due to their adaptation to extreme environmental niche conditions. However, NmCyp exhibits low thermal stability and starts to aggregate beyond 40 °C. The properties of NmCyp are compared to those reported for the cyclophilin from another archaeal organism, Methanobrevibacter ruminantium. The current study sheds light on the differential behavior of cyclophilin proteins from two different phyla of archaea.
© 2021 The Authors. Published by American Chemical Society.
Conflict of interest statement
The authors declare no competing financial interest.
Figures


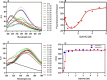
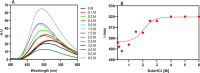
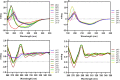
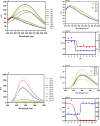
References
LinkOut - more resources
Full Text Sources

