The Role of Antibiotic Resistance Genes in the Fitness Cost of Multiresistance Plasmids
- PMID: 35038907
- PMCID: PMC8764527
- DOI: 10.1128/mbio.03552-21
The Role of Antibiotic Resistance Genes in the Fitness Cost of Multiresistance Plasmids
Abstract
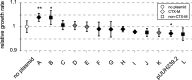
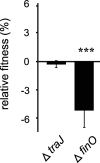
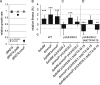
By providing the bacterial cell with protection against several antibiotics at once, multiresistance plasmids have an evolutionary advantage in situations where antibiotic treatments are common, such as in hospital environments. However, resistance plasmids can also impose fitness costs on the bacterium in the absence of antibiotics, something that may limit their evolutionary success. The underlying mechanisms and the possible contribution of resistance genes to such costs are still largely not understood. Here, we have specifically investigated the contribution of plasmid-borne resistance genes to the reduced fitness of the bacterial cell. The pUUH239.2 plasmid carries 13 genes linked to antibiotic resistance and reduces bacterial fitness by 2.9% per generation. This cost is fully ameliorated by the removal of the resistance cassette. While most of the plasmid-borne resistance genes individually were cost-free, even when overexpressed, two specific gene clusters were responsible for the entire cost of the plasmid: the extended-spectrum-β-lactamase gene blaCTX-M-15 and the tetracycline resistance determinants tetAR. The blaCTX-M-15 cost was linked to the signal peptide that exports the β-lactamase into the periplasm, and replacement with an alternative signal peptide abolished the cost. Both the tetracycline pump TetA and its repressor TetR conferred a cost on the host cell, and the reciprocal expression of these genes is likely fine-tuned to balance the respective costs. These findings highlight that the cost of clinical multiresistance plasmids can be largely due to particular resistance genes and their interaction with other cellular systems, while other resistance genes and the plasmid backbone can be cost-free. IMPORTANCE Multiresistance plasmids are one of the main drivers of antibiotic resistance development and spread. Their evolutionary success through the accumulation and mobilization of resistance genes is central to resistance evolution. In this study, we find that the cost of the introduction of a multiresistance plasmid was completely attributable to resistance genes, while the rest of the plasmid backbone is cost-free. The majority of resistance genes on the plasmid had no appreciable cost to the host cell even when overexpressed, indicating that plasmid-borne resistance can be cost-free. In contrast, the widespread genes blaCTX-M-15 and tetAR were found to confer the whole cost of the plasmid by affecting specific cellular functions. These findings highlight how the evolution of resistance on plasmids is dependent on the amelioration of associated fitness costs and point at a conundrum regarding the high cost of some of the most widespread β-lactamase genes.
Keywords: antibiotic resistance; fitness cost; plasmid-mediated resistance.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

