Functional characterisation of the amyotrophic lateral sclerosis risk locus GPX3/TNIP1
- PMID: 35042540
- PMCID: PMC8767698
- DOI: 10.1186/s13073-021-01006-6
Functional characterisation of the amyotrophic lateral sclerosis risk locus GPX3/TNIP1
Abstract
Background: Amyotrophic lateral sclerosis (ALS) is a complex, late-onset, neurodegenerative disease with a genetic contribution to disease liability. Genome-wide association studies (GWAS) have identified ten risk loci to date, including the TNIP1/GPX3 locus on chromosome five. Given association analysis data alone cannot determine the most plausible risk gene for this locus, we undertook a comprehensive suite of in silico, in vivo and in vitro studies to address this.
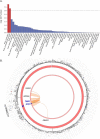
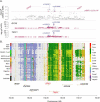
Methods: The Functional Mapping and Annotation (FUMA) pipeline and five tools (conditional and joint analysis (GCTA-COJO), Stratified Linkage Disequilibrium Score Regression (S-LDSC), Polygenic Priority Scoring (PoPS), Summary-based Mendelian Randomisation (SMR-HEIDI) and transcriptome-wide association study (TWAS) analyses) were used to perform bioinformatic integration of GWAS data (Ncases = 20,806, Ncontrols = 59,804) with 'omics reference datasets including the blood (eQTLgen consortium N = 31,684) and brain (N = 2581). This was followed up by specific expression studies in ALS case-control cohorts (microarray Ntotal = 942, protein Ntotal = 300) and gene knockdown (KD) studies of human neuronal iPSC cells and zebrafish-morpholinos (MO).
Results: SMR analyses implicated both TNIP1 and GPX3 (p < 1.15 × 10-6), but there was no simple SNP/expression relationship. Integrating multiple datasets using PoPS supported GPX3 but not TNIP1. In vivo expression analyses from blood in ALS cases identified that lower GPX3 expression correlated with a more progressed disease (ALS functional rating score, p = 5.5 × 10-3, adjusted R2 = 0.042, Beffect = 27.4 ± 13.3 ng/ml/ALSFRS unit) with microarray and protein data suggesting lower expression with risk allele (recessive model p = 0.06, p = 0.02 respectively). Validation in vivo indicated gpx3 KD caused significant motor deficits in zebrafish-MO (mean difference vs. control ± 95% CI, vs. control, swim distance = 112 ± 28 mm, time = 1.29 ± 0.59 s, speed = 32.0 ± 2.53 mm/s, respectively, p for all < 0.0001), which were rescued with gpx3 expression, with no phenotype identified with tnip1 KD or gpx3 overexpression.
Conclusions: These results support GPX3 as a lead ALS risk gene in this locus, with more data needed to confirm/reject a role for TNIP1. This has implications for understanding disease mechanisms (GPX3 acts in the same pathway as SOD1, a well-established ALS-associated gene) and identifying new therapeutic approaches. Few previous examples of in-depth investigations of risk loci in ALS exist and a similar approach could be applied to investigate future expected GWAS findings.
Keywords: Computational biology; Disease progression; Genes; Genome-wide association study; MND; Motor neurone disease; Neurodegenerative diseases; Quantitative trait loci; Regulator; Zebrafish.
© 2021. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures




References
-
- Benyamin B, He J, Zhao Q, Gratten J, Garton F, Leo PJ, Liu Z, Mangelsdorf M, al-Chalabi A, Anderson L, Butler TJ, Chen L, Chen XD, Cremin K, Deng HW, Devine M, Edson J, Fifita JA, Furlong S, Han YY, Harris J, Henders AK, Jeffree RL, Jin ZB, Li Z, Li T, Li M, Lin Y, Liu X, Marshall M, McCann EP, Mowry BJ, Ngo ST, Pamphlett R, Ran S, Reutens DC, Rowe DB, Sachdev P, Shah S, Song S, Tan LJ, Tang L, van den Berg LH, van Rheenen W, Veldink JH, Wallace RH, Wheeler L, Williams KL, Wu J, Wu X, Yang J, Yue W, Zhang ZH, Zhang D, Noakes PG, Blair IP, Henderson RD, McCombe PA, Visscher PM, Xu H, Bartlett PF, Brown MA, Wray NR, Fan D. Cross-ethnic meta-analysis identifies association of the GPX3-TNIP1 locus with amyotrophic lateral sclerosis. Nat Commun. 2017;8(1):611. doi: 10.1038/s41467-017-00471-1. - DOI - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

