The antigenicity of SARS-CoV-2 Delta variants aggregated 10 high-frequency mutations in RBD has not changed sufficiently to replace the current vaccine strain
- PMID: 35046385
- PMCID: PMC8767530
- DOI: 10.1038/s41392-022-00874-7
The antigenicity of SARS-CoV-2 Delta variants aggregated 10 high-frequency mutations in RBD has not changed sufficiently to replace the current vaccine strain
Abstract
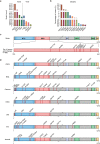
Emerging SARS-CoV-2 variants are the most serious problem for COVID-19 prophylaxis and treatment. To determine whether the SARS-CoV-2 vaccine strain should be updated following variant emergence like seasonal flu vaccine, the changed degree on antigenicity of SARS-CoV-2 variants and H3N2 flu vaccine strains was compared. The neutralization activities of Alpha, Beta and Gamma variants' spike protein-immunized sera were analysed against the eight current epidemic variants and 20 possible variants combining the top 10 prevalent RBD mutations based on the Delta variant, which were constructed using pseudotyped viruses. Meanwhile, the neutralization activities of convalescent sera and current inactivated and recombinant protein vaccine-elicited sera were also examined against all possible Delta variants. Eight HA protein-expressing DNAs elicited-animal sera were also tested against eight pseudotyped viruses of H3N2 flu vaccine strains from 2011-2019. Our results indicate that the antigenicity changes of possible Delta variants were mostly within four folds, whereas the antigenicity changes among different H3N2 vaccine strains were approximately 10-100-fold. Structural analysis of the antigenic characterization of the SARS-CoV-2 and H3N2 mutations supports the neutralization results. This study indicates that the antigenicity changes of the current SARS-CoV-2 may not be sufficient to require replacement of the current vaccine strain.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures




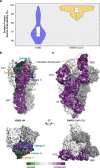
References
Publication types
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

