Mechanisms of Polycomb group protein function in cancer
- PMID: 35046519
- PMCID: PMC8888700
- DOI: 10.1038/s41422-021-00606-6
Mechanisms of Polycomb group protein function in cancer
Abstract
Cancer arises from a multitude of disorders resulting in loss of differentiation and a stem cell-like phenotype characterized by uncontrolled growth. Polycomb Group (PcG) proteins are members of multiprotein complexes that are highly conserved throughout evolution. Historically, they have been described as essential for maintaining epigenetic cellular memory by locking homeotic genes in a transcriptionally repressed state. What was initially thought to be a function restricted to a few target genes, subsequently turned out to be of much broader relevance, since the main role of PcG complexes is to ensure a dynamically choregraphed spatio-temporal regulation of their numerous target genes during development. Their ability to modify chromatin landscapes and refine the expression of master genes controlling major switches in cellular decisions under physiological conditions is often misregulated in tumors. Surprisingly, their functional implication in the initiation and progression of cancer may be either dependent on Polycomb complexes, or specific for a subunit that acts independently of other PcG members. In this review, we describe how misregulated Polycomb proteins play a pleiotropic role in cancer by altering a broad spectrum of biological processes such as the proliferation-differentiation balance, metabolism and the immune response, all of which are crucial in tumor progression. We also illustrate how interfering with PcG functions can provide a powerful strategy to counter tumor progression.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures

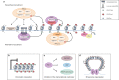
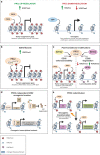

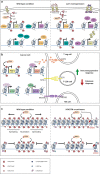
References
-
- Loubiere V, Martinez AM, Cavalli G. Cell fate and developmental regulation dynamics by polycomb proteins and 3D genome architecture. BioEssays. 2019;41:1–15. - PubMed
-
- Chan HL, Morey L. Emerging roles for polycomb-group proteins in stem cells and cancer. Trends Biochem. Sci. 2019;44:688–700. - PubMed
-
- Piunti A, Shilatifard A. The roles of polycomb repressive complexes in mammalian development and cancer. Nat. Rev. Mol. Cell Biol. 2021;22:326–345. - PubMed
-
- Lewis EB. A gene complex controlling segmentation in Drosophila. Nature. 1978;276:565–570. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

