Pyruvate dehydrogenase inactivation causes glycolytic phenotype in BAP1 mutant uveal melanoma
- PMID: 35046531
- PMCID: PMC9066178
- DOI: 10.1038/s41388-021-02154-0
Pyruvate dehydrogenase inactivation causes glycolytic phenotype in BAP1 mutant uveal melanoma
Abstract
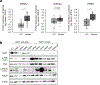
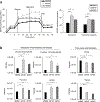
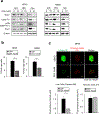
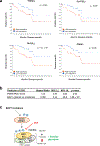
Effective therapeutic options are still lacking for uveal melanoma (UM) patients who develop metastasis. Metastatic traits of UM are linked to BRCA1-associated protein 1 (BAP1) mutations. Cell metabolism is re-programmed in UM with BAP1 mutant UM, but the underlying mechanisms and opportunities for therapeutic intervention remain unclear. BAP1 mutant UM tumors have an elevated glycolytic gene expression signature, with increased expression of pyruvate dehydrogenase (PDH) complex and PDH kinase (PDHK1). Furthermore, BAP1 mutant UM cells showed higher levels of phosphorylated PDHK1 and PDH that was associated with an upregulated glycolytic profile compared to BAP1 wild-type UM cells. Suppressing PDHK1-PDH phosphorylation decreased glycolytic capacity and cell growth, and induced cell cycle arrest of BAP1 mutant UM cells. Our results suggest that PDHK1-PDH phosphorylation is a causative factor of glycolytic phenotypes found in BAP1 mutant UM and propose a therapeutic opportunity for BAP1 mutant UM patients.
© 2021. The Author(s), under exclusive licence to Springer Nature Limited.
Conflict of interest statement
Figures



References
-
- Singh N, Bergman L, Seregard S, Singh AD. Uveal melanoma: Epidemiologic aspects. Ocul Oncol Pathol: Springer; 2014. p. 75–87.
-
- Chua V, Aplin AE. Novel therapeutic strategies and targets in advanced uveal melanoma. Curr Opin Oncol. 2018;30(2):134–41. - PubMed
-
- Carvajal RD, Piperno-Neumann S, Kapiteijn E, Chapman PB, Frank S, Joshua AM, et al. Selumetinib in combination with dacarbazine in patients with metastatic uveal melanoma: a phase III, multicentre, randomised trial (SUMIT). J Clin Oncol. 2018;36(12):1232–9. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

