A Workflow of Integrated Resources to Catalyze Network Pharmacology Driven COVID-19 Research
- PMID: 35057621
- PMCID: PMC10790216
- DOI: 10.1021/acs.jcim.1c00431
A Workflow of Integrated Resources to Catalyze Network Pharmacology Driven COVID-19 Research
Abstract
In the event of an outbreak due to an emerging pathogen, time is of the essence to contain or to mitigate the spread of the disease. Drug repositioning is one of the strategies that has the potential to deliver therapeutics relatively quickly. The SARS-CoV-2 pandemic has shown that integrating critical data resources to drive drug-repositioning studies, involving host-host, host-pathogen, and drug-target interactions, remains a time-consuming effort that translates to a delay in the development and delivery of a life-saving therapy. Here, we describe a workflow we designed for a semiautomated integration of rapidly emerging data sets that can be generally adopted in a broad network pharmacology research setting. The workflow was used to construct a COVID-19 focused multimodal network that integrates 487 host-pathogen, 63 278 host-host protein, and 1221 drug-target interactions. The resultant Neo4j graph database named "Neo4COVID19" is made publicly accessible via a web interface and via API calls based on the Bolt protocol. Details for accessing the database are provided on a landing page (https://neo4covid19.ncats.io/). We believe that our Neo4COVID19 database will be a valuable asset to the research community and will catalyze the discovery of therapeutics to fight COVID-19.
Figures
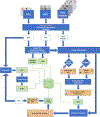


Update of
-
A Workflow of Integrated Resources to Catalyze Network Pharmacology Driven COVID-19 Research.bioRxiv [Preprint]. 2020 Nov 5:2020.11.04.369041. doi: 10.1101/2020.11.04.369041. bioRxiv. 2020. Update in: J Chem Inf Model. 2022 Feb 14;62(3):718-729. doi: 10.1021/acs.jcim.1c00431. PMID: 33173863 Free PMC article. Updated. Preprint.
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

