Genome-wide methylation analyses identifies Non-coding RNA genes dysregulated in breast tumours that metastasise to the brain
- PMID: 35058523
- PMCID: PMC8776809
- DOI: 10.1038/s41598-022-05050-z
Genome-wide methylation analyses identifies Non-coding RNA genes dysregulated in breast tumours that metastasise to the brain
Abstract
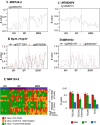
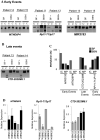
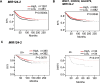
Brain metastases comprise 40% of all metastatic tumours and breast tumours are among the tumours that most commonly metastasise to the brain, the role that epigenetic gene dysregulation plays in this process is not well understood. We carried out 450 K methylation array analysis to investigate epigenetically dysregulated genes in breast to brain metastases (BBM) compared to normal breast tissues (BN) and primary breast tumours (BP). For this, we referenced 450 K methylation data for BBM tumours prepared in our laboratory with BN and BP from The Cancer Genome Atlas. Experimental validation on our initially identified genes, in an independent cohort of BP and in BBM and their originating primary breast tumours using Combined Bisulphite and Restriction Analysis (CoBRA) and Methylation Specific PCR identified three genes (RP11-713P17.4, MIR124-2, NUS1P3) that are hypermethylated and three genes (MIR3193, CTD-2023M8.1 and MTND6P4) that are hypomethylated in breast to brain metastases. In addition, methylation differences in candidate genes between BBM tumours and originating primary tumours shows dysregulation of DNA methylation occurs either at an early stage of tumour evolution (in the primary tumour) or at a later evolutionary stage (where the epigenetic change is only observed in the brain metastasis). Epigentic changes identified could also be found when analysing tumour free circulating DNA (tfcDNA) in patient's serum taken during BBM biopsies. Epigenetic dysregulation of RP11-713P17.4, MIR3193, MTND6P4 are early events suggesting a potential use for these genes as prognostic markers.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures


Similar articles
-
The GALNT9, BNC1 and CCDC8 genes are frequently epigenetically dysregulated in breast tumours that metastasise to the brain.Clin Epigenetics. 2015 May 27;7(1):57. doi: 10.1186/s13148-015-0089-x. eCollection 2015. Clin Epigenetics. 2015. PMID: 26052355 Free PMC article.
-
An integrated epigenomic-transcriptomic landscape of lung cancer reveals novel methylation driver genes of diagnostic and therapeutic relevance.Theranostics. 2021 Mar 11;11(11):5346-5364. doi: 10.7150/thno.58385. eCollection 2021. Theranostics. 2021. PMID: 33859751 Free PMC article.
-
Exploring breast carcinogenesis through integrative genomics and epigenomics analyses.Int J Oncol. 2014 Nov;45(5):1959-68. doi: 10.3892/ijo.2014.2625. Epub 2014 Aug 27. Int J Oncol. 2014. PMID: 25175708
-
Epigenetics of Circulating Tumor Cells in Breast Cancer.Adv Exp Med Biol. 2020;1220:117-134. doi: 10.1007/978-3-030-35805-1_8. Adv Exp Med Biol. 2020. PMID: 32304083 Review.
-
Epigenetic Biomarkers of Breast Cancer Risk: Across the Breast Cancer Prevention Continuum.Adv Exp Med Biol. 2016;882:33-68. doi: 10.1007/978-3-319-22909-6_2. Adv Exp Med Biol. 2016. PMID: 26987530 Free PMC article. Review.
Cited by
-
Insights into the role of long non-coding RNAs in DNA methylation mediated transcriptional regulation.Front Mol Biosci. 2022 Dec 2;9:1067406. doi: 10.3389/fmolb.2022.1067406. eCollection 2022. Front Mol Biosci. 2022. PMID: 36533073 Free PMC article. Review.
-
Advances in Brain Metastases Diagnosis: Non-coding RNAs As Potential Biomarkers.Cureus. 2023 Mar 18;15(3):e36337. doi: 10.7759/cureus.36337. eCollection 2023 Mar. Cureus. 2023. PMID: 37077610 Free PMC article. Review.
-
Metastatic brain tumors: from development to cutting-edge treatment.MedComm (2020). 2024 Dec 20;6(1):e70020. doi: 10.1002/mco2.70020. eCollection 2025 Jan. MedComm (2020). 2024. PMID: 39712454 Free PMC article. Review.
-
The Role of Methylation as an Epigenetic Marker in HPV-Related Oral Lesions.J Med Virol. 2025 Jul;97(7):e70459. doi: 10.1002/jmv.70459. J Med Virol. 2025. PMID: 40556436 Free PMC article.
-
Liquid biopsies to occult brain metastasis.Mol Cancer. 2022 May 10;21(1):113. doi: 10.1186/s12943-022-01577-x. Mol Cancer. 2022. PMID: 35538484 Free PMC article. Review.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

