MinION™ Nanopore Sequencing of Skin Microbiome 16S and 16S-23S rRNA Gene Amplicons
- PMID: 35071053
- PMCID: PMC8766866
- DOI: 10.3389/fcimb.2021.806476
MinION™ Nanopore Sequencing of Skin Microbiome 16S and 16S-23S rRNA Gene Amplicons
Abstract
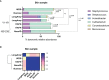
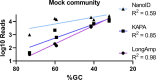
Human skin microbiome dysbiosis can have clinical consequences. Characterizing taxonomic composition of bacterial communities associated with skin disorders is important for dermatological advancement in both diagnosis and novel treatments. This study aims to analyze and improve the accuracy of taxonomic classification of skin bacteria with MinION™ nanopore sequencing using a defined skin mock community and a skin microbiome sample. We compared the Oxford Nanopore Technologies recommended procedures and concluded that their protocols highly bias the relative abundance of certain skin microbiome genera, most notably a large overrepresentation of Staphylococcus and underrepresentation of Cutibacterium and Corynebacterium. We demonstrated that changes in the amplification protocols improved the accuracy of the taxonomic classification for these three main skin bacterial genera. This study shows that MinION™ nanopore could be an efficient technology for full-length 16S rRNA sequencing; however, the analytical advantage is strongly influenced by the methodologies. The suggested alternatives in the sample processing improved characterization of a complex skin microbiome community using MinION™ nanopore sequencing.
Keywords: 16S rRNA gene sequencing; MinION™; bacterial identification; nanopore sequencing; skin microbiome; skin mock community.
Copyright © 2022 Rozas, Brillet, Callewaert and Paetzold.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

