Clonally expanded B cells in multiple sclerosis bind EBV EBNA1 and GlialCAM
- PMID: 35073561
- PMCID: PMC9382663
- DOI: 10.1038/s41586-022-04432-7
Clonally expanded B cells in multiple sclerosis bind EBV EBNA1 and GlialCAM
Abstract
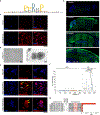
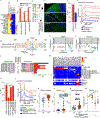
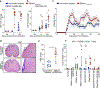
Multiple sclerosis (MS) is a heterogenous autoimmune disease in which autoreactive lymphocytes attack the myelin sheath of the central nervous system. B lymphocytes in the cerebrospinal fluid (CSF) of patients with MS contribute to inflammation and secrete oligoclonal immunoglobulins1,2. Epstein-Barr virus (EBV) infection has been epidemiologically linked to MS, but its pathological role remains unclear3. Here we demonstrate high-affinity molecular mimicry between the EBV transcription factor EBV nuclear antigen 1 (EBNA1) and the central nervous system protein glial cell adhesion molecule (GlialCAM) and provide structural and in vivo functional evidence for its relevance. A cross-reactive CSF-derived antibody was initially identified by single-cell sequencing of the paired-chain B cell repertoire of MS blood and CSF, followed by protein microarray-based testing of recombinantly expressed CSF-derived antibodies against MS-associated viruses. Sequence analysis, affinity measurements and the crystal structure of the EBNA1-peptide epitope in complex with the autoreactive Fab fragment enabled tracking of the development of the naive EBNA1-restricted antibody to a mature EBNA1-GlialCAM cross-reactive antibody. Molecular mimicry is facilitated by a post-translational modification of GlialCAM. EBNA1 immunization exacerbates disease in a mouse model of MS, and anti-EBNA1 and anti-GlialCAM antibodies are prevalent in patients with MS. Our results provide a mechanistic link for the association between MS and EBV and could guide the development of new MS therapies.
© 2022. This is a U.S. government work and not under copyright protection in the U.S.; foreign copyright protection may apply.
Conflict of interest statement
Figures










Comment in
-
EBV linked to multiple sclerosis.Nat Rev Microbiol. 2022 Apr;20(4):189. doi: 10.1038/s41579-022-00701-4. Nat Rev Microbiol. 2022. PMID: 35110729 No abstract available.
-
Linking Epstein-Barr virus infection to multiple sclerosis.Nat Rev Immunol. 2022 Mar;22(3):143. doi: 10.1038/s41577-022-00686-4. Nat Rev Immunol. 2022. PMID: 35140366 No abstract available.
-
Epstein-Barr virus sparks brain autoimmunity in multiple sclerosis.Nature. 2022 Mar;603(7900):230-232. doi: 10.1038/d41586-022-00382-2. Nature. 2022. PMID: 35169323 No abstract available.
-
Guilty by association: Epstein-Barr virus in multiple sclerosis.Nat Med. 2022 May;28(5):904-906. doi: 10.1038/s41591-022-01823-1. Nat Med. 2022. PMID: 35538259 No abstract available.
-
Targeting Epstein-Barr virus to treat MS.Med. 2022 Mar 11;3(3):159-161. doi: 10.1016/j.medj.2022.02.005. Med. 2022. PMID: 35590190
-
A role for the Epstein-Barr virus in multiple sclerosis aetiology?J Neurol. 2022 Jul;269(7):3962-3963. doi: 10.1007/s00415-022-11177-w. Epub 2022 May 31. J Neurol. 2022. PMID: 35639199 Free PMC article. No abstract available.
References
-
- Cencioni MT, Mattoscio M, Magliozzi R, Bar-Or A & Muraro PA B cells in multiple sclerosis - from targeted depletion to immune reconstitution therapies. Nat. Rev. Neurol. 17, 399–414 (2021). - PubMed
-
- Hauser SL et al. Ocrelizumab versus Interferon Beta-1a in Relapsing Multiple Sclerosis. N. Engl. J. Med. 376, 221–234 (2017). - PubMed
-
- Jarius S et al. The MRZ reaction as a highly specific marker of multiple sclerosis: re-evaluation and structured review of the literature. J. Neurol. 264, 453–466 (2017). - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

