Colon stroma mediates an inflammation-driven fibroblastic response controlling matrix remodeling and healing
- PMID: 35085231
- PMCID: PMC8824371
- DOI: 10.1371/journal.pbio.3001532
Colon stroma mediates an inflammation-driven fibroblastic response controlling matrix remodeling and healing
Abstract
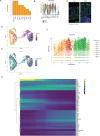
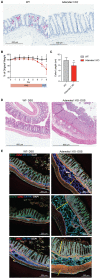
Chronic inflammation is often associated with the development of tissue fibrosis, but how mesenchymal cell responses dictate pathological fibrosis versus resolution and healing remains unclear. Defining stromal heterogeneity and identifying molecular circuits driving extracellular matrix deposition and remodeling stands to illuminate the relationship between inflammation, fibrosis, and healing. We performed single-cell RNA-sequencing of colon-derived stromal cells and identified distinct classes of fibroblasts with gene signatures that are differentially regulated by chronic inflammation, including IL-11-producing inflammatory fibroblasts. We further identify a transcriptional program associated with trans-differentiation of mucosa-associated fibroblasts and define a functional gene signature associated with matrix deposition and remodeling in the inflamed colon. Our analysis supports a critical role for the metalloprotease Adamdec1 at the interface between tissue remodeling and healing during colitis, demonstrating its requirement for colon epithelial integrity. These findings provide mechanistic insight into how inflammation perturbs stromal cell behaviors to drive fibroblastic responses controlling mucosal matrix remodeling and healing.
Conflict of interest statement
I have read the journal’s policy and the authors of this manuscript have the following competing interests: R.J.X. is a cofounder of Celsius Therapeutics and Jnana Therapeutics.
Figures




References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

