Variability in Host Specificity and Functional Potential of Antarctic Sponge-Associated Bacterial Communities
- PMID: 35095792
- PMCID: PMC8792898
- DOI: 10.3389/fmicb.2021.771589
Variability in Host Specificity and Functional Potential of Antarctic Sponge-Associated Bacterial Communities
Abstract
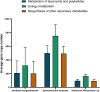
Sponge-associated microorganisms are essential for sponge survival. They play an important role in recycling nutrients and, therefore, in the maintenance of the ecosystem. These microorganisms are diverse, species-specific, and different from those in the surrounding seawater. Bacterial sponge symbionts have been extensively studied in the tropics; however, little is known about these microorganisms in sponges from high-latitude environments. Sponges can cover up to 80% of the benthos in Antarctica and are crucial architects for the marine food web. In this study, we present analyses of the bacterial symbionts of three sponges: Haliclona (Rhizoniera) sp., Hymeniacidon torquata, and Isodictya kerguelenensis from the Western Antarctic Peninsula (WAP) with the aim to determine variations on the specificity of the bacteria-sponge interactions and potential signatures on their predicted functional profiles. We use high-throughput 16S rRNA gene sequencing of 30 sponge individuals inhabiting South Bay (Palmer Archipelago, WAP) to describe their microbiome taxonomy and diversity and predict potential functional profiles based on this marker gene. Our work shows similar bacterial community composition profiles among the same sponge species, although the symbiotic relationship is not equally conserved among the three Antarctic sponges. The number of species-specific core operational taxonomic units (OTUs) of these Antarctic sponges was low, with important differences between the total abundance accounted for these OTUs. Only eight OTUs were shared between the three sponge species. Analyses of the functional potential revealed that despite the high host-symbiont specificity, the inferred functions are conserved among these microbiomes, although with differences in the abundance of specific functions. H. torquata showed the highest level of intra-specificity and a higher potential of pathways related to energy metabolism, metabolisms of terpenoids and polyketides, and biosynthesis of other secondary metabolites. Overall, this work shows variations in the specificity of the sponge-associated bacterial communities, differences in how hosts and symbionts establish their relations, and in their potential functional capabilities.
Keywords: 16S rRNA gene; Antarctic sponges; functional potential; high-throughput sequencing; host specificity; microbiome; secondary metabolites; symbiosis.
Copyright © 2022 Cristi, Parada-Pozo, Morales-Vicencio, Cárdenas and Trefault.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
Similar articles
-
Temporal Stability of Bacterial Communities in Antarctic Sponges.Front Microbiol. 2019 Nov 22;10:2699. doi: 10.3389/fmicb.2019.02699. eCollection 2019. Front Microbiol. 2019. PMID: 31824467 Free PMC article.
-
Characterization of Bacterial, Archaeal and Eukaryote Symbionts from Antarctic Sponges Reveals a High Diversity at a Three-Domain Level and a Particular Signature for This Ecosystem.PLoS One. 2015 Sep 30;10(9):e0138837. doi: 10.1371/journal.pone.0138837. eCollection 2015. PLoS One. 2015. PMID: 26421612 Free PMC article.
-
Evidence of habitat specificity in sponge microbiomes from Antarctica.Environ Microbiome. 2024 Dec 4;19(1):100. doi: 10.1186/s40793-024-00648-4. Environ Microbiome. 2024. PMID: 39633476 Free PMC article.
-
The Microbial Ecology of Antarctic Sponges.Microb Ecol. 2025 May 17;88(1):44. doi: 10.1007/s00248-025-02543-y. Microb Ecol. 2025. PMID: 40382475 Free PMC article. Review.
-
Comparative Metagenomic Analysis of Biosynthetic Diversity across Sponge Microbiomes Highlights Metabolic Novelty, Conservation, and Diversification.mSystems. 2022 Aug 30;7(4):e0035722. doi: 10.1128/msystems.00357-22. Epub 2022 Jul 18. mSystems. 2022. PMID: 35862823 Free PMC article. Review.
Cited by
-
Bacteria Associated with Benthic Invertebrates from Extreme Marine Environments: Promising but Underexplored Sources of Biotechnologically Relevant Molecules.Mar Drugs. 2022 Sep 29;20(10):617. doi: 10.3390/md20100617. Mar Drugs. 2022. PMID: 36286440 Free PMC article. Review.
-
Current knowledge of the Southern Hemisphere marine microbiome in eukaryotic hosts and the Strait of Magellan surface microbiome project.PeerJ. 2023 Oct 3;11:e15978. doi: 10.7717/peerj.15978. eCollection 2023. PeerJ. 2023. PMID: 37810788 Free PMC article. Review.
-
A comparison of free-living and sponge-associated bacterial communities from a remote oceanic island with a focus on calcareous sponges.FEMS Microbiol Ecol. 2023 Feb 28;99(3):fiad014. doi: 10.1093/femsec/fiad014. FEMS Microbiol Ecol. 2023. PMID: 36758964 Free PMC article.
-
Chemical and microbiological insights into two littoral Antarctic demosponge species: Haliclona (Rhizoniera) dancoi (Topsent 1901) and Haliclona (Rhizoniera) scotti (Kirkpatrick 1907).Front Microbiol. 2024 Feb 9;15:1341641. doi: 10.3389/fmicb.2024.1341641. eCollection 2024. Front Microbiol. 2024. PMID: 38404594 Free PMC article.
-
Stability of the Microbiome of the Sponge Mycale (Oxymycale) acerata in the Western Antarctic Peninsula.Front Microbiol. 2022 Apr 4;13:827863. doi: 10.3389/fmicb.2022.827863. eCollection 2022. Front Microbiol. 2022. PMID: 35444618 Free PMC article.
References
-
- Barnett D., Arts I., Penders J. (2021). microViz: an R package for microbiome data visualization and statistics. J. Open Source Softw. 6:3201. 10.21105/joss.03201 - DOI
-
- Bell J. J. (2008). The functional roles of marine sponges. Estuar. Coast. Shelf Sci. 79 341–353. 10.1016/j.ecss.2008.05.002 - DOI
-
- Berne S., Kalauz M., Lapat M., Savin L., Janussen D., Kersken D., et al. (2016). Screening of the Antarctic marine sponges (Porifera) as a source of bioactive compounds. Polar Biol. 39 947–959. 10.1007/s00300-015-1835-4 - DOI
LinkOut - more resources
Full Text Sources

