The FUT2 Variant c.461G>A (p.Trp154*) Is Associated With Differentially Expressed Genes and Nasopharyngeal Microbiota Shifts in Patients With Otitis Media
- PMID: 35096646
- PMCID: PMC8798324
- DOI: 10.3389/fcimb.2021.798246
The FUT2 Variant c.461G>A (p.Trp154*) Is Associated With Differentially Expressed Genes and Nasopharyngeal Microbiota Shifts in Patients With Otitis Media
Abstract
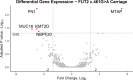
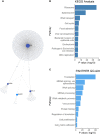
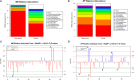
Otitis media (OM) is a leading cause of childhood hearing loss. Variants in FUT2, which encodes alpha-(1,2)-fucosyltransferase, were identified to increase susceptibility to OM, potentially through shifts in the middle ear (ME) or nasopharyngeal (NP) microbiotas as mediated by transcriptional changes. Greater knowledge of differences in relative abundance of otopathogens in carriers of pathogenic variants can help determine risk for OM in patients. In order to determine the downstream effects of FUT2 variation, we examined gene expression in relation to carriage of a common pathogenic FUT2 c.461G>A (p.Trp154*) variant using RNA-sequence data from saliva samples from 28 patients with OM. Differential gene expression was also examined in bulk mRNA and single-cell RNA-sequence data from wildtype mouse ME mucosa after inoculation with non-typeable Haemophilus influenzae (NTHi). In addition, microbiotas were profiled from ME and NP samples of 65 OM patients using 16S rRNA gene sequencing. In human carriers of the FUT2 variant, FN1, KMT2D, MUC16 and NBPF20 were downregulated while MTAP was upregulated. Post-infectious expression in the mouse ME recapitulated these transcriptional differences, with the exception of Fn1 upregulation after NTHi-inoculation. In the NP, Candidate Division TM7 was associated with wildtype genotype (FDR-adj-p=0.009). Overall, the FUT2 c.461G>A variant was associated with transcriptional changes in processes related to response to infection and with increased load of potential otopathogens in the ME and decreased commensals in the NP. These findings provide increased understanding of how FUT2 variants influence gene transcription and the mucosal microbiota, and thus contribute to the pathology of OM.
Keywords: FUT2; RNA-sequencing; microbiota; otitis media; p.Trp154*; rs601338.
Copyright © 2022 Elling, Scholes, Streubel, Larson, Wine, Bootpetch, Yoon, Kofonow, Gubbels, Cass, Robertson, Jenkins, Prager, Frank, Chan, Friedman, Ryan and Santos-Cortez.
Conflict of interest statement
AR is a cofounder of Otonomy, Inc., serves as a consultant and member of the Scientific Advisory Board, and holds an equity position in the company. The UCSD and San Diego VA Committees on Conflict of Interest have approved this relationship. Otonomy, Inc. had no role in the conduct of this study and the writing of the manuscript. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures




References
-
- Brennan-Jones C. G., Whitehouse A. J., Park J., Hegarty M., Jacques A., Eikelboom R. H., et al. . (2015). Prevalence and Risk Factors for Parent-Reported Recurrent Otitis Media During Early Childhood in the Western Australian Pregnancy Cohort (Raine) Study. J. Paediatr. Child Health 51 (4), 403–409. doi: 10.1111/jpc.12741 - DOI - PubMed
-
- Browne J. J., Matthews E. H., Taylor-Robinson A. W., Kyd J. M. (2021). Otopathogen Interactions in the Nasopharynx of Children, and the Predictive Value of Nasopharyngeal Aspirate Culture for the Aetiology of Upper Respiratory Infections. J. Paediatr. Child Health 57 (7), 1016–1022. doi: 10.1111/jpc.15370 - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

