Vimentin binds to G-quadruplex repeats found at telomeres and gene promoters
- PMID: 35100428
- PMCID: PMC8860586
- DOI: 10.1093/nar/gkab1274
Vimentin binds to G-quadruplex repeats found at telomeres and gene promoters
Abstract
G-quadruplex (G4) structures that can form at guanine-rich genomic sites, including telomeres and gene promoters, are actively involved in genome maintenance, replication, and transcription, through finely tuned interactions with protein networks. In the present study, we identified the intermediate filament protein Vimentin as a binder with nanomolar affinity for those G-rich sequences that give rise to at least two adjacent G4 units, named G4 repeats. This interaction is supported by the N-terminal domains of soluble Vimentin tetramers. The selectivity of Vimentin for G4 repeats versus individual G4s provides an unprecedented result. Based on GO enrichment analysis performed on genes having putative G4 repeats within their core promoters, we suggest that Vimentin recruitment at these sites may contribute to the regulation of gene expression during cell development and migration, possibly by reshaping the local higher-order genome topology, as already reported for lamin B.
© The Author(s) 2022. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures
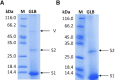

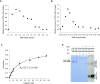
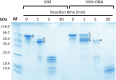
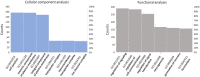
Similar articles
-
Interaction in vitro of type III intermediate filament proteins with higher order structures of single-stranded DNA, particularly with G-quadruplex DNA.DNA Cell Biol. 2005 Feb;24(2):85-110. doi: 10.1089/dna.2005.24.85. DNA Cell Biol. 2005. PMID: 15699629
-
Non-duplex G-Quadruplex DNA Structure: A Developing Story from Predicted Sequences to DNA Structure-Dependent Epigenetics and Beyond.Acc Chem Res. 2021 Jan 5;54(1):46-56. doi: 10.1021/acs.accounts.0c00431. Epub 2020 Dec 21. Acc Chem Res. 2021. PMID: 33347280 Free PMC article.
-
G-quadruplex-R-loop interactions and the mechanism of anticancer G-quadruplex binders.Nucleic Acids Res. 2020 Dec 2;48(21):11942-11957. doi: 10.1093/nar/gkaa944. Nucleic Acids Res. 2020. PMID: 33137181 Free PMC article. Review.
-
G-quadruplexes formation within the promoter of TEAD4 oncogene and their interaction with Vimentin.Front Chem. 2022 Sep 15;10:1008075. doi: 10.3389/fchem.2022.1008075. eCollection 2022. Front Chem. 2022. PMID: 36186582 Free PMC article.
-
G-Quadruplex-Binding Proteins: Promising Targets for Drug Design.Biomolecules. 2022 Apr 29;12(5):648. doi: 10.3390/biom12050648. Biomolecules. 2022. PMID: 35625576 Free PMC article. Review.
Cited by
-
Roles of G4-DNA and G4-RNA in Class Switch Recombination and Additional Regulations in B-Lymphocytes.Molecules. 2023 Jan 24;28(3):1159. doi: 10.3390/molecules28031159. Molecules. 2023. PMID: 36770824 Free PMC article. Review.
-
Polymorphic and Higher-Order G-Quadruplexes as Possible Transcription Regulators: Novel Perspectives for Future Anticancer Therapeutic Applications.Pharmaceuticals (Basel). 2022 Mar 19;15(3):373. doi: 10.3390/ph15030373. Pharmaceuticals (Basel). 2022. PMID: 35337170 Free PMC article.
-
Vimentin: from a cytoskeletal protein to a critical modulator of immune response and a target for infection.Front Immunol. 2023 Jul 5;14:1224352. doi: 10.3389/fimmu.2023.1224352. eCollection 2023. Front Immunol. 2023. PMID: 37475865 Free PMC article. Review.
-
Vimentin Is at the Heart of Epithelial Mesenchymal Transition (EMT) Mediated Metastasis.Cancers (Basel). 2021 Oct 5;13(19):4985. doi: 10.3390/cancers13194985. Cancers (Basel). 2021. PMID: 34638469 Free PMC article. Review.
-
Higher-order G-quadruplexes in promoters are untapped drug targets.Front Chem. 2023 Jun 7;11:1211512. doi: 10.3389/fchem.2023.1211512. eCollection 2023. Front Chem. 2023. PMID: 37351517 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

