Plasma tRNA-derived small RNAs signature as a predictive and prognostic biomarker in lung adenocarcinoma
- PMID: 35115004
- PMCID: PMC8812260
- DOI: 10.1186/s12935-022-02481-6
Plasma tRNA-derived small RNAs signature as a predictive and prognostic biomarker in lung adenocarcinoma
Abstract
Background: The prevalence of lung adenocarcinoma (LUAD) has increased, thus novel biomarkers for its early diagnosis is becoming more important than ever. tRNA-derived small RNA (tsRNA) is a new class of non-coding RNA which has important regulatory roles in cancer biology. This study was designed to identify novel predictive and prognostic tsRNA biomarkers.
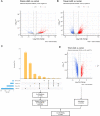
Methods: tsRNAs were identified and performed differential expression analysis from 10 plasma samples (6 LUAD and 4 normal, SRP266333) and 96 tissue samples (48 LUAD and 48 normal, SRP133217). Then a tsRNA-mRNA regulatory network was constructed to find hub tsRNAs. Functional enrichment analysis was performed to infer the potential pathways associated with tsRNAs. Afterwards, a Support Vector Machine (SVM) algorithm was used to explore the potential biomarkers for diagnosing LUAD. Lastly, the function of tRF-21-RK9P4P9L0 was explored in A549 and H1299 cell lines.
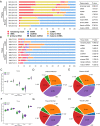
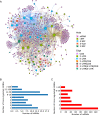
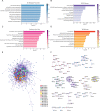
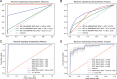
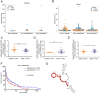
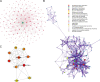
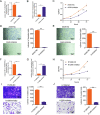
Results: A significant difference of read distribution was observed between normal people and LUAD patients whether in plasma or tissue. A tsRNA-mRNA regulatory network consisting of 155 DEtsRNAs (differential expression tsRNAs) and 406 DEmRNAs (differential expression mRNAs) was established. Three tsRNAs (tRF-16-L85J3KE, tRF-21-RK9P4P9L0 and tRF-16-PSQP4PE) were identified as hub genes with degree > 100. We found Co-DEmRNAs (intersection of DEtsRNAs target mRNAs and differentially expressed mRNAs in LUAD) were engaged in a number of cancer pathways. The AUC of the three hub tsRNAs' expression for diagnosing LUAD reached 0.92. Furthermore, the qPCR validation of the three hub tsRNAs in 37 paired normal and LUAD tissues was consistent with the RNA-Seq results. In addition, tRF-21-RK9P4P9L0 was negatively associated with LUAD prognosis. Inhibition of tRF-21-RK9P4P9L0 expression reduced the proliferation, migration and invasion ability of A549 and H1299 cell lines.
Conclusion: These findings will help us further understand the molecular mechanisms of LUAD and contribute to novel diagnostic biomarkers and therapeutic target discovery.
Keywords: Diagnostic biomarker; Lung adenocarcinoma; Network; tRFs; tsRNAs.
© 2022. The Author(s).
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures
Similar articles
-
Serum tRNA-derived small RNAs as potential novel diagnostic biomarkers for pancreatic ductal adenocarcinoma.Am J Cancer Res. 2021 Mar 1;11(3):837-848. eCollection 2021. Am J Cancer Res. 2021. PMID: 33791157 Free PMC article.
-
Transfer RNA-derived small RNAs (tsRNAs) sequencing revealed a differential expression landscape of tsRNAs between glioblastoma and low-grade glioma.Gene. 2023 Mar 1;855:147114. doi: 10.1016/j.gene.2022.147114. Epub 2022 Dec 13. Gene. 2023. PMID: 36526122
-
Global profile of tRNA-derived small RNAs in gastric cancer patient plasma and identification of tRF-33-P4R8YP9LON4VDP as a new tumor suppressor.Int J Med Sci. 2021 Feb 4;18(7):1570-1579. doi: 10.7150/ijms.53220. eCollection 2021. Int J Med Sci. 2021. PMID: 33746573 Free PMC article.
-
The regulatory role of tRNA-derived small RNAs in the prognosis of gastric cancer.Cell Signal. 2025 Jan;125:111511. doi: 10.1016/j.cellsig.2024.111511. Epub 2024 Nov 16. Cell Signal. 2025. PMID: 39551416 Review.
-
Unveiling the role of tRNA-derived small RNAs in MAPK signaling pathway: implications for cancer and beyond.Front Genet. 2024 Mar 26;15:1346852. doi: 10.3389/fgene.2024.1346852. eCollection 2024. Front Genet. 2024. PMID: 38596214 Free PMC article. Review.
Cited by
-
Diagnostic and prognostic values of tsRNAs in lung cancer: a meta-analysis.BMC Cancer. 2025 Jan 27;25(1):153. doi: 10.1186/s12885-025-13536-y. BMC Cancer. 2025. PMID: 39871144 Free PMC article.
-
The role and mechanism of action of tRNA-derived fragments in the diagnosis and treatment of malignant tumors.Cell Commun Signal. 2023 Mar 24;21(1):62. doi: 10.1186/s12964-023-01079-3. Cell Commun Signal. 2023. PMID: 36964534 Free PMC article. Review.
-
Multiple regulatory roles of the transfer RNA-derived small RNAs in cancers.Genes Dis. 2023 Apr 13;11(2):597-613. doi: 10.1016/j.gendis.2023.02.053. eCollection 2024 Mar. Genes Dis. 2023. PMID: 37692525 Free PMC article. Review.
-
tsRNA: A Promising Biomarker in Breast Cancer.J Cancer. 2024 Mar 11;15(9):2613-2626. doi: 10.7150/jca.93531. eCollection 2024. J Cancer. 2024. PMID: 38577588 Free PMC article. Review.
-
The biological role and molecular mechanism of transfer RNA-derived small RNAs in tumor metastasis.Front Oncol. 2025 Apr 8;15:1560943. doi: 10.3389/fonc.2025.1560943. eCollection 2025. Front Oncol. 2025. PMID: 40265011 Free PMC article. Review.
References
-
- Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424. - PubMed
LinkOut - more resources
Full Text Sources

