RNA kink-turns are highly anisotropic with respect to lateral displacement of the flanking stems
- PMID: 35122735
- PMCID: PMC8943727
- DOI: 10.1016/j.bpj.2022.01.025
RNA kink-turns are highly anisotropic with respect to lateral displacement of the flanking stems
Erratum in
-
RNA kink-turns are highly anisotropic with respect to lateral displacement of the flanking stems.Biophys J. 2025 Jul 1;124(13):2251. doi: 10.1016/j.bpj.2025.05.024. Epub 2025 May 28. Biophys J. 2025. PMID: 40441138 Free PMC article. No abstract available.
Abstract
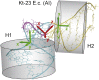
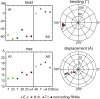
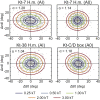
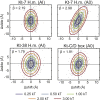
Kink-turns are highly bent internal loop motifs commonly found in the ribosome and other RNA complexes. They frequently act as binding sites for proteins and mediate tertiary interactions in larger RNA structures. Kink-turns have been a topic of intense research, but their elastic properties in the folded state are still poorly understood. Here we use extensive all-atom molecular dynamics simulations to parameterize a model of kink-turn in which the two flanking helical stems are represented by effective rigid bodies. Time series of the full set of six interhelical coordinates enable us to extract minimum energy shapes and harmonic stiffness constants for kink-turns from different RNA functional classes. The analysis suggests that kink-turns exhibit isotropic bending stiffness but are highly anisotropic with respect to lateral displacement of the stems. The most flexible lateral displacement mode is perpendicular to the plane of the static bend. These results may help understand the structural adaptation and mechanical signal transmission by kink-turns in complex natural and artificial RNA structures.
Copyright © 2022 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Figures




References
-
- Ben-Shem A., Garreau de Loubresse N., et al. Yusupov M. The structure of the eukaryotic ribosome at 3.0 A resolution. Science. 2011;334:1524–1529. - PubMed
-
- Moore T., Zhang Y., et al. Li H. Molecular basis of box C/D RNA-protein interactions: cocrystal structure of Archaeal L7Ae and a box C/D RNA. Structure. 2004;12:807–818. - PubMed
-
- Montange R.K., Batey R.T. Structure of the S-adenosylmethionine riboswitch regulatory mRNA element. Nature. 2006;441:1172–1175. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

