Metagenomics of the midgut microbiome of Rhipicephalus microplus from China
- PMID: 35135613
- PMCID: PMC8822867
- DOI: 10.1186/s13071-022-05161-6
Metagenomics of the midgut microbiome of Rhipicephalus microplus from China
Abstract
Background: Ticks, which are ectoparasites of animals, may carry multiple pathogens. The cattle tick Rhipicephalus microplus is an important bovine parasite in China. However, the midgut microbiome of R. microplus from China has not been characterized via metagenomic methods.
Methods: Rhipicephalus microplus were collected from cattle in the city of Changsha in Hunan province, China. The DNA of the midgut contents was extracted from fully engorged adult female R. microplus. A DNA library was constructed and sequenced using an Illumina HiSeq sequencing platform. SOAPdenovo software was used to assemble and analyze the clean data. The latent class analysis algorithm applied to system classification by MEGAN software was used to annotate the information on the species' sequences. DIAMOND software was used to compare unigenes with the Kyoto Encyclopedia of Genes and Genomes (KEGG) database, and functional annotation was carried out based on the results of the comparison.
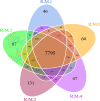
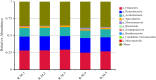
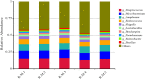
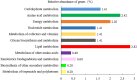
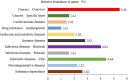
Results: The dominant phyla in the five samples were Firmicutes, Proteobacteria, and Actinobacteria. Streptococcus, Mycobacterium, Anaplasma, Enterococcus, Shigella, Lactobacillus, Brachyspira, Pseudomonas, Enterobacter, Bacillus, and Lactococcus were the dominant genera in the five samples. The endosymbiotic bacterium Wolbachia was also detected in all of the samples. Mycobacterium malmesburyense, Streptococcus pneumoniae, Anaplasma phagocytophilum, Enterococcus faecium, Shigella sonnei, Enterococcus faecalis, Lactobacillus casei, Brachyspira hampsonii, Pseudomonas syringae, Enterobacter cloacae, and Lactococcus garvieae were the dominant species in the five samples. In addition to these bacterial species, we also detected some eukaryotes, such as Rhizophagus irregularis, Enterospora canceri, Smittium culicis, Zancudomyces culisetae, Trachipleistophora hominis, and viruses such as orf virus, human endogenous retrovirus type W, enzootic nasal tumor virus of goats, bovine retrovirus CH15, and galidia endogenous retrovirus in all of the samples at the species level. The results of the annotated KEGG pathway predictions for the gene functions of the midgut microflora of R. microplus indicated genes involved in lipid and amino acid metabolism, infectious diseases (e.g., Streptococcus pneumonia infection, human granulocytic anaplasmosis, Shigella sonnei infection, Salmonella enterica infection, and pathogenic Escherichia coli infection), and cancer.
Conclusions: Our study revealed that the midgut microbiome of R. microplus is not only composed of a large number of bacteria, but that a portion also comprises eukaryotes and viruses. The data presented here enhance our understanding of this tick's midgut microbiome and provide fundamental information for the control of ticks and tick-borne diseases.
Keywords: Gene function; Metagenomics; Microbiome; Midgut; Rhipicephalus microplus.
© 2022. The Author(s).
Conflict of interest statement
The authors declare that they have no conflicts of interest.
Figures


Similar articles
-
Metagenome reveals the midgut microbial community of Haemaphysalis qinghaiensis ticks collected from yaks and Tibetan sheep.Parasit Vectors. 2024 Aug 31;17(1):370. doi: 10.1186/s13071-024-06442-y. Parasit Vectors. 2024. PMID: 39217389 Free PMC article.
-
Prevalence, risk factors, and genetic diversity of veterinary important tick-borne pathogens in cattle from Rhipicephalus microplus-invaded and non-invaded areas of Benin.Ticks Tick Borne Dis. 2018 Mar;9(3):450-464. doi: 10.1016/j.ttbdis.2017.12.015. Epub 2017 Dec 28. Ticks Tick Borne Dis. 2018. PMID: 29307783
-
Comparative microarray analyses of adult female midgut tissues from feeding Rhipicephalus species.Ticks Tick Borne Dis. 2015 Feb;6(1):84-90. doi: 10.1016/j.ttbdis.2014.09.008. Epub 2014 Oct 22. Ticks Tick Borne Dis. 2015. PMID: 25448423
-
A review of the history of research and control of Rhipicephalus (Boophilus) microplus, babesiosis and anaplasmosis in Uruguay.Exp Appl Acarol. 2018 Aug;75(4):383-398. doi: 10.1007/s10493-018-0278-3. Epub 2018 Aug 6. Exp Appl Acarol. 2018. PMID: 30083875 Review.
-
Review of cattle ticks (Acari, Ixodida) in Ivory Coast and geographic distribution of Rhipicephalus (Boophilus) microplus, an emerging tick in West Africa.Exp Appl Acarol. 2017 Apr;71(4):355-369. doi: 10.1007/s10493-017-0129-7. Epub 2017 May 11. Exp Appl Acarol. 2017. PMID: 28497303 Review.
Cited by
-
Microbial agents for the control of ticks Rhipicephalus microplus.Parasitol Res. 2024 Jul 17;123(7):275. doi: 10.1007/s00436-024-08291-1. Parasitol Res. 2024. PMID: 39017922 Review.
-
Metagenome reveals the midgut microbial community of Haemaphysalis qinghaiensis ticks collected from yaks and Tibetan sheep.Parasit Vectors. 2024 Aug 31;17(1):370. doi: 10.1186/s13071-024-06442-y. Parasit Vectors. 2024. PMID: 39217389 Free PMC article.
-
Characterization and manipulation of the bacterial community in the midgut of Ixodes ricinus.Parasit Vectors. 2022 Jul 9;15(1):248. doi: 10.1186/s13071-022-05362-z. Parasit Vectors. 2022. PMID: 35810301 Free PMC article.
-
Diversity of Rickettsiales in Rhipicephalus microplus Ticks Collected in Domestic Ruminants in Guizhou Province, China.Pathogens. 2022 Sep 27;11(10):1108. doi: 10.3390/pathogens11101108. Pathogens. 2022. PMID: 36297165 Free PMC article.
-
Genomic assessment of targets implicated in Rhipicephalus microplus acaricide resistance.PLoS One. 2024 Dec 5;19(12):e0312074. doi: 10.1371/journal.pone.0312074. eCollection 2024. PLoS One. 2024. PMID: 39637189 Free PMC article.
References
-
- de Barros MNDL, Riet-Correa F, Azevedo SS, Labruna MB. Off-host development and survival of Rhipicephalus (Boophilus) microplus in the Brazilian semiarid. Vet Parasitol Reg Stud Reports. 2017;9:17–24. - PubMed
-
- Vesco U, Knap N, Labruna MB, Avsic-Zupanc T, Estrada-Pena A, Guglielmone AA, et al. An integrated database on ticks and tick-borne zoonoses in the tropics and subtropics with special reference to developing and emerging countries. Exp Appl Acarol. 2011;54:65–83. - PubMed
-
- Estrada-Peña A, García Z, Sánchez HF. The distribution and ecological preferences of Boophilus microplus (Acari: Ixodidae) in Mexico. Exp Appl Acarol. 2006;38:307–316. - PubMed
-
- Nava S, Mastropaolo M, Guglielmone AA, Mangold AJ. Effect of deforestation and introduction of exotic grasses as livestock forage on the population dynamics of the cattle tick Rhipicephalus (Boophilus) microplus (Acari: Ixodidae) in northern Argentina. Res Vet Sci. 2013;95:1046–1054. - PubMed
-
- Chen Z, Yang X, Bu F, Yang X, Yang X, Liu J. Ticks (acari: ixodoidea: argasidae, ixodidae) of China. Exp Appl Acarol. 2010;51:393–404. - PubMed
MeSH terms
Grants and funding
- 31902294/the National Science Foundation of China
- 2018RS3085/the Planned Programme of Hunan Province Science and Technology Innovation
- KH2002001/the Training Programme for Excellent Young Innovators of Changsha
- 2020JJ5230/the Natural Science Foundation of Hunan Province, China
- 19A235/the Research Foundation of Education Bureau of Hunan Province, China
LinkOut - more resources
Full Text Sources
Medical

