Chlamydomonas CHT7 is involved in repressing DNA replication and mitotic genes during synchronous growth
- PMID: 35137070
- PMCID: PMC8895990
- DOI: 10.1093/g3journal/jkac023
Chlamydomonas CHT7 is involved in repressing DNA replication and mitotic genes during synchronous growth
Abstract
In the green alga Chlamydomonas reinhardtii, regulation of the cell cycle in response to external cues is critical for survival in a changing environment. The loss of the nuclear COMPROMISED HYDROLYSIS OF TRIACYLGLYCEROLS 7 (CHT7) protein affects the expression of many genes especially in response to nitrogen availability. Cells lacking CHT7 exhibit abnormal cell morphology following nitrogen deprivation and fail to resume normal cell division after N resupply. To investigate the function of CHT7 in the regulation of cell cycle-related pathways, cells were synchronized, and RNA-seq analysis was performed during various stages of the cell cycle. In the cht7 mutant following nitrogen deprivation, the cells were not dividing, but a subset of cell cycle genes involved in DNA replication and mitosis were found to be derepressed, suggesting that the CHT7 protein plays a role in cell cycle regulation that is opposite to that of the mitotic cyclin-dependent kinases. Furthermore, genes for cell wall synthesis and remodeling were found to be abnormally induced in nondividing cht7 cells; this misregulation may deplete cellular resources and thus contribute to cell death following nitrogen deprivation. Lastly, 43 minimally characterized kinases were found to be highly misregulated in cht7. Further analysis suggested that some of these CHT7-regulated kinases may be related to the MAP3K and Aurora-like kinases, while others are unique. Together, these results suggest a role of CHT7 in transcriptional regulation of the cell cycle and reveal several pathways and genes whose expression appears to be subject to a CHT7-mediated regulatory network.
Keywords: Chlamydomonas reinhardtii; DNA replication; algal cell wall; cell cycle; cell division; mitosis; quiescence.
© The Author(s) 2022. Published by Oxford University Press on behalf of Genetics Society of America.
Figures


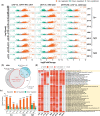





Similar articles
-
Chlamydomonas CHT7 Is Required for an Effective Quiescent State by Regulating Nutrient-Responsive Cell Cycle Gene Expression.Plant Cell. 2020 Apr;32(4):1240-1269. doi: 10.1105/tpc.19.00628. Epub 2020 Jan 30. Plant Cell. 2020. PMID: 32001503 Free PMC article.
-
Modulation of CHT7 Complexes during Light/Dark- and Nitrogen-Mediated Life Cycle Transitions of Chlamydomonas.Plant Physiol. 2020 Dec;184(4):1762-1774. doi: 10.1104/pp.20.00864. Epub 2020 Oct 1. Plant Physiol. 2020. PMID: 33004613 Free PMC article.
-
The protein Compromised Hydrolysis of Triacylglycerols 7 (CHT7) acts as a repressor of cellular quiescence in Chlamydomonas.Proc Natl Acad Sci U S A. 2014 Nov 4;111(44):15833-8. doi: 10.1073/pnas.1414567111. Epub 2014 Oct 13. Proc Natl Acad Sci U S A. 2014. PMID: 25313078 Free PMC article.
-
Nitrogen-dependent coordination of cell cycle, quiescence and TAG accumulation in Chlamydomonas.Biotechnol Biofuels. 2019 Dec 23;12:292. doi: 10.1186/s13068-019-1635-0. eCollection 2019. Biotechnol Biofuels. 2019. PMID: 31890020 Free PMC article. Review.
-
The Chlamydomonas cell cycle.Plant J. 2015 May;82(3):370-392. doi: 10.1111/tpj.12795. Epub 2015 Apr 15. Plant J. 2015. PMID: 25690512 Free PMC article. Review.
Cited by
-
The micromammals.G3 (Bethesda). 2024 Jun 5;14(6):jkae073. doi: 10.1093/g3journal/jkae073. G3 (Bethesda). 2024. PMID: 38837137 Free PMC article.
References
-
- Alexa A, Rahnenfuhrer J. topGO: Enrichment Analysis for Gene Ontology. R Package Version 2.42.0; 2020.
-
- Alexa A, Rahnenfuhrer J, Lengauer T.. Improved scoring of functional groups from gene expression data by decorrelating GO graph structure. Bioinformatics. 2006;22(13):1600–1607. - PubMed
-
- Anisimova M, Gascuel O.. Approximate likelihood-ratio test for branches: a fast, accurate, and powerful alternative. Syst Biol. 2006;55(4):539–552. - PubMed
-
- Beall EL, Manak JR, Zhou S, Bell M, Lipsick JS, Botchan MR.. Role for a Drosophila Myb-containing protein complex in site-specific DNA replication. Nature. 2002;420(6917):833–837. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
