Genetic Determinants of Ammonium Excretion in nifL Mutants of Azotobacter vinelandii
- PMID: 35138932
- PMCID: PMC8939361
- DOI: 10.1128/AEM.01876-21
Genetic Determinants of Ammonium Excretion in nifL Mutants of Azotobacter vinelandii
Abstract
The ubiquitous diazotrophic soil bacterium Azotobacter vinelandii has been extensively studied as a model organism for biological nitrogen fixation (BNF). In A. vinelandii, BNF is regulated by the NifL-NifA two-component system, where NifL acts as an antiactivator that tightly controls the activity of the nitrogen fixation-specific transcriptional activator NifA in response to redox, nitrogen, and carbon status. While several studies reported that mutations in A. vinelandii nifL resulted in the deregulation of nitrogenase expression and the release of large quantities of ammonium, knowledge about the specific determinants for this ammonium-excreting phenotype is lacking. In this work, we report that only specific disruptions of nifL lead to large quantities of ammonium accumulated in liquid culture (∼12 mM). The ammonium excretion phenotype is associated solely with deletions of NifL domains combined with the insertion of a promoter sequence in the orientation opposite that of nifLA transcription. We further demonstrated that the strength of the inserted promoter could influence the amounts of ammonium excreted by affecting rnf1 gene expression as an additional requirement for ammonium excretion. These ammonium-excreting nifL mutants significantly stimulate the transfer of fixed nitrogen to rice. This work defines discrete determinants that bring about A. vinelandii ammonium excretion and demonstrates that strains can be generated through simple gene editing to provide promising biofertilizers capable of transferring nitrogen to crops. IMPORTANCE There is considerable interest in the engineering of ammonium-excreting bacteria for use in agriculture to promote the growth of plants under fixed-nitrogen-limiting conditions. This work defines discrete determinants that bring about A. vinelandii ammonium excretion and demonstrates that strains can be generated through simple gene editing to provide promising biofertilizers capable of transferring nitrogen to crops.
Keywords: A. vinelandii; ammonium excretion; biofertilizer; nifL; nitrogen fixation; regulation; rice; transfer of fixed nitrogen.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
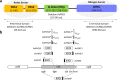
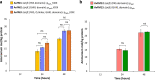
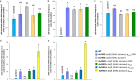
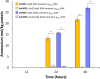
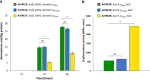
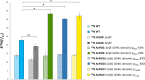
References
-
- Borlaug NE. 1997. Feeding a world of 10 billion people: the miracle ahead. Biotechnol Dynamo Equip 11:3–13. 10.1080/13102818.1997.10818934. - DOI
-
- Erisman JW, Galloway JN, Dise NB, Sutton MA, Bleeker A, Grizzetti B, Leach AM, De Vries W. 2015. Nitrogen: too much of a vital resource. WWF science brief NL. WWF Netherlands, Zeist, The Netherlands.
-
- Mus F, Crook MB, Garcia K, Garcia Costas A, Geddes BA, Kouri ED, Paramasivan P, Ryu M-H, Oldroyd GED, Poole PS, Udvardi MK, Voigt CA, Ané J-M, Peters JW. 2016. Symbiotic nitrogen fixation and the challenges to its extension to nonlegumes. Appl Environ Microbiol 82:3698–3710. 10.1128/AEM.01055-16. - DOI - PMC - PubMed
-
- Colnaghi R, Green A, He L, Rudnick P, Kennedy C. 1997. Strategies for increased ammonium production in free-living or plant associated nitrogen fixing bacteria. Plant Soil 194:145–154. 10.1023/A:1004268526162. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

