Nrf2 Inhibits the Progression of Parkinson's Disease by Upregulating AABR07032261.5 to Repress Pyroptosis
- PMID: 35140498
- PMCID: PMC8818975
- DOI: 10.2147/JIR.S345895
Nrf2 Inhibits the Progression of Parkinson's Disease by Upregulating AABR07032261.5 to Repress Pyroptosis
Abstract
Objective: Parkinson's disease (PD) is associated with dysregulated neural cell death, such as pyroptosis, but its regulatory mechanisms are poorly understood. This study investigated roles of nuclear factor E2-related factor 2 (Nrf2) in regulating pyroptosis and PD development.
Methods: Cellular and rat PD models established by 6-OHDA exposure were subjected to Nrf2 overexpression. Neurobehavioral functions were assessed by the traction test, Morris Water Maze, and open field test. Cell proliferation was analyzed by MTS assay, while flow cytometry was applied to quantify levels of reactive oxygen species (ROS) and apoptosis. Nissl bodies in rat brains were detected by Nissl staining, and cell apoptosis in brain tissues was assessed by terminal deoxynucleotidyl transferase dUTP nick-end labeling. Differential expression of lncRNA and mRNA was characterized by deep sequencing.
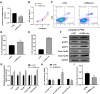
Results: A cellular PD model was successfully established by inducing PC12 cell differentiation with nerve growth factor-β and exposing differentiated cells to 6-OHDA. Cells exhibited significantly increased ROS levels, enhanced pyroptosis, and inhibited Nrf2 phosphorylation. The rat PD model exhibited impaired muscle strength, increased pyroptosis, and repressed Nrf2 phosphorylation. Nrf2 overexpression effectively repressed pyroptosis in both cellular and rat PD models. Marked alterations of lncRNA and mRNA profiles were induced by Nrf2 overexpression in the cellular PD model, which involved multiple signaling pathways. Silencing of the lncRNA AABR07032261.5 significantly promoted pyroptosis in the cellular PD model.
Conclusion: Nrf2 suppressed PD pathogenesis in cellular and animal models by promoting AABR07032261.5, which repressed pyroptosis.
Keywords: AABR07032261.5; Nrf2; Parkinson’s disease; lncRNAs; oxidative stress; pyroptosis.
© 2022 Zhong et al.
Conflict of interest statement
The authors report no conflicts of interest in this work.
Figures





Similar articles
-
Chlorpyrifos activates cell pyroptosis and increases susceptibility on oxidative stress-induced toxicity by miR-181/SIRT1/PGC-1α/Nrf2 signaling pathway in human neuroblastoma SH-SY5Y cells: Implication for association between chlorpyrifos and Parkinson's disease.Environ Toxicol. 2019 Jun;34(6):699-707. doi: 10.1002/tox.22736. Epub 2019 Mar 5. Environ Toxicol. 2019. PMID: 30835941
-
Inhibition of PRMT5 Attenuates Oxidative Stress-Induced Pyroptosis via Activation of the Nrf2/HO-1 Signal Pathway in a Mouse Model of Renal Ischemia-Reperfusion Injury.Oxid Med Cell Longev. 2019 Nov 25;2019:2345658. doi: 10.1155/2019/2345658. eCollection 2019. Oxid Med Cell Longev. 2019. PMID: 31885778 Free PMC article.
-
LncRNA MALAT1 facilitates inflammasome activation via epigenetic suppression of Nrf2 in Parkinson's disease.Mol Brain. 2020 Sep 24;13(1):130. doi: 10.1186/s13041-020-00656-8. Mol Brain. 2020. PMID: 32972446 Free PMC article.
-
Long non-coding RNA myocardial infarction-associated transcript promotes 1-Methyl-4-phenylpyridinium ion-induced neuronal inflammation and oxidative stress in Parkinson's disease through regulating microRNA-221-3p/ transforming growth factor /nuclear factor E2-related factor 2 axis.Bioengineered. 2022 Jan;13(1):930-940. doi: 10.1080/21655979.2021.2015527. Bioengineered. 2022. PMID: 34967706 Free PMC article.
-
Dimethyl fumarate attenuates 6-OHDA-induced neurotoxicity in SH-SY5Y cells and in animal model of Parkinson's disease by enhancing Nrf2 activity.Neuroscience. 2015 Feb 12;286:131-40. doi: 10.1016/j.neuroscience.2014.11.047. Epub 2014 Nov 29. Neuroscience. 2015. PMID: 25449120
Cited by
-
Triple-Combination Therapy with a Multifunctional Yolk-Shell Nanozyme Au@CeO2 Loaded with Dimethyl Fumarate for Periodontitis.Adv Sci (Weinh). 2025 Feb;12(7):e2413891. doi: 10.1002/advs.202413891. Epub 2024 Dec 24. Adv Sci (Weinh). 2025. PMID: 39716921 Free PMC article.
-
Chronic Cerebral Hypoperfusion Aggravates Parkinson's Disease Dementia-Like Symptoms and Pathology in 6-OHDA-Lesioned Rat through Interfering with Sphingolipid Metabolism.Oxid Med Cell Longev. 2022 Aug 8;2022:5392966. doi: 10.1155/2022/5392966. eCollection 2022. Oxid Med Cell Longev. 2022. PMID: 35979400 Free PMC article.
-
Possible Mitigating Effect of Nobiletin on Rotenone-Induced Parkinsonism in Male Albino Rat Model: Targeting Bmal1/Nrf2 Mediated Ferroptosis and Restoring Mitochondrial Mitophagy.Neurochem Res. 2025 Jun 25;50(4):208. doi: 10.1007/s11064-025-04438-3. Neurochem Res. 2025. PMID: 40563018
-
The Role of Astrocytes and Alpha-Synuclein in Parkinson's Disease: A Review.NeuroSci. 2024 Mar 8;5(1):71-86. doi: 10.3390/neurosci5010005. eCollection 2024 Mar. NeuroSci. 2024. PMID: 39483813 Free PMC article. Review.
-
Parkinson's disease models and death signaling: what do we know until now?Front Neuroanat. 2024 Oct 29;18:1419108. doi: 10.3389/fnana.2024.1419108. eCollection 2024. Front Neuroanat. 2024. PMID: 39533977 Free PMC article. Review.
References
LinkOut - more resources
Full Text Sources

