Immune cell gene expression signatures in diffuse glioma are associated with IDH mutation status, patient outcome and malignant cell state, and highlight the importance of specific cell subsets in glioma biology
- PMID: 35144680
- PMCID: PMC8830123
- DOI: 10.1186/s40478-022-01323-w
Immune cell gene expression signatures in diffuse glioma are associated with IDH mutation status, patient outcome and malignant cell state, and highlight the importance of specific cell subsets in glioma biology
Abstract
The tumor micro-environment (TME) plays an important role in various cancers, including gliomas. We estimated immune cell type-specific gene expression profiles in 3 large clinically annotated glioma datasets using CIBERSORTx and LM22/LM10 blood-based immune signatures and found that the proportions and estimated gene expression patterns of specific immune cells significantly varied according to IDH mutation status. When IDH-WT and IDH-MUT tumors were considered separately, cluster-of-cluster analyses of immune cell gene expression identified groups with distinct survival outcomes. We confirmed and extended these findings by applying a signature matrix derived from single-cell RNA-sequencing data derived from 19 glioma tumor samples to the bulk profiling data, validating findings from the LM22/LM10 results. To link immune cell signatures with outcomes in checkpoint therapy, we then showed a significant association of monocytic lineage cell gene expression clusters with patient survival and with mesenchymal gene expression scores. Integrating immune cell-based gene expression with previously described malignant cell states in glioma demonstrated that macrophage M0 abundance significantly correlated with mesenchymal state in IDH-WT gliomas, with evidence of a previously implicated role of the Oncostatin-M receptor and macrophages in the mesenchymal state. Among IDH-WT tumors that were enriched for the mesenchymal cell state, the estimated M0 macrophage expression signature coordinately also trended to a mesenchymal signature. We also examined IDH-MUT tumors stratified by 1p/19q status, showing that a mesenchymal gene expression signature the M0 macrophage fraction was enriched in IDH-MUT, non-codeleted tumors. Overall, these results highlight the biological and clinical significance of the immune cell environment related to IDH mutation status, patient prognosis and the mesenchymal state in diffuse gliomas.
Keywords: CIBERSORTx; Deconvolution; Glioma; Malignant cell-state; Prognosis; Tumor microenvironment.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures


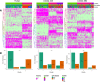




References
-
- Ascierto ML, Kmieciak M, Idowu MO, Manjili R, Zhao Y, Grimes M, Dumur C, Wang E, Ramakrishnan V, Wang XY, et al. A signature of immune function genes associated with recurrence-free survival in breast cancer patients. Breast Cancer Res Treat. 2012;131:871–880. doi: 10.1007/s10549-011-1470-x. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

